6.4.6. Feature Importance#
Your Random Forest achieves 95% accuracy predicting customer churn. But why? Which features matter most?
Feature importance tells us:
Which features the model relies on
Which features are redundant
What patterns the model found
Why it matters:
Interpretability: Explain to stakeholders
Feature selection: Remove unimportant features
Debug: Detect data leakage or bad features
Trust: Verify model makes sense
Let’s explore different methods to understand what models learn!
6.4.6.1. The Fundamental Question#
Question: If I remove or shuffle a feature, how much does performance drop?
High importance → Performance drops significantly Low importance → Performance barely affected
6.4.6.2. Built-In Feature Importance (Tree Models)#
Tree-based models have built-in feature importance!
How it works:*
Importance = How much each feature decreases impurity
Accumulated across all trees in forest
Normalized to sum to 1
Pros: Fast, automatic Cons: Biased toward high-cardinality features
cancer = load_breast_cancer()
X_cancer, y_cancer = cancer.data, cancer.target
feature_names = cancer.feature_names
X_train, X_test, y_train, y_test = train_test_split(
X_cancer, y_cancer, test_size=0.3, random_state=42, stratify=y_cancer
)
rf = RandomForestClassifier(n_estimators=100, random_state=42)
rf.fit(X_train, y_train)
glue('rf-acc', round(rf.score(X_test, y_test), 3), display=False)
importances = rf.feature_importances_
indices = np.argsort(importances)[::-1]
cumsum_importances = np.cumsum(importances[indices])
glue('top5-cumulative', f'{cumsum_importances[4]:.1%}', display=False)
glue('top10-cumulative', f'{cumsum_importances[9]:.1%}', display=False)
top10_df = pd.DataFrame([
{'Rank': i+1, 'Feature': feature_names[indices[i]], 'Importance': round(importances[indices[i]], 4)}
for i in range(10)
])
display(top10_df)
fig, axes = plt.subplots(1, 2, figsize=(14, 6))
top_10_idx = indices[:10]
axes[0].barh(range(10), importances[top_10_idx], alpha=0.7, edgecolor='black')
axes[0].set_yticks(range(10))
axes[0].set_yticklabels([feature_names[i] for i in top_10_idx], fontsize=9)
axes[0].set_xlabel('Importance', fontsize=12)
axes[0].set_title('Top 10 Most Important Features\n(Random Forest)', fontsize=13, fontweight='bold')
axes[0].invert_yaxis()
axes[0].grid(True, alpha=0.3, axis='x')
axes[1].barh(range(len(importances)), importances[indices], alpha=0.7, edgecolor='black')
axes[1].set_xlabel('Importance', fontsize=12)
axes[1].set_ylabel('Feature Index (sorted)', fontsize=12)
axes[1].set_title('All Features Importance Distribution\n(Long tail of unimportant features)', fontsize=13, fontweight='bold')
axes[1].grid(True, alpha=0.3, axis='x')
plt.tight_layout()
plt.show()
| Rank | Feature | Importance | |
|---|---|---|---|
| 0 | 1 | worst concave points | 0.1590 |
| 1 | 2 | worst area | 0.1470 |
| 2 | 3 | worst perimeter | 0.0858 |
| 3 | 4 | worst radius | 0.0790 |
| 4 | 5 | mean radius | 0.0777 |
| 5 | 6 | mean perimeter | 0.0742 |
| 6 | 7 | mean concave points | 0.0659 |
| 7 | 8 | mean concavity | 0.0543 |
| 8 | 9 | mean area | 0.0417 |
| 9 | 10 | worst concavity | 0.0314 |
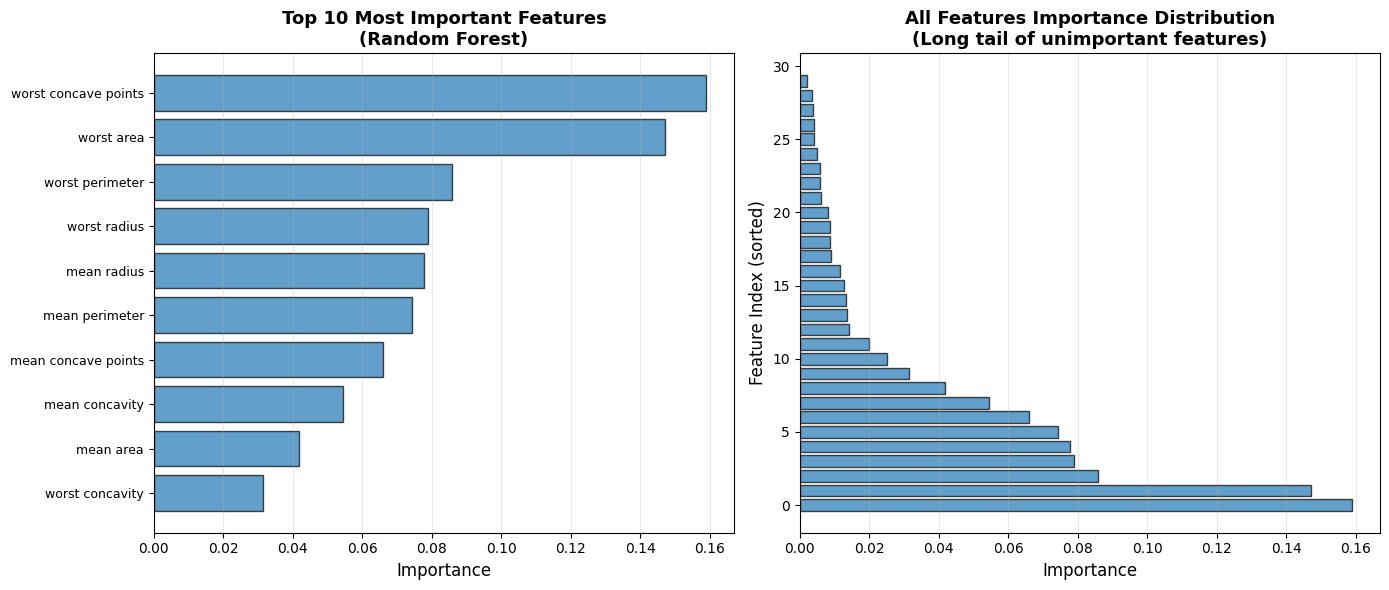
Random Forest accuracy: 0.936. The top 5 features account for 54.8% of total importance; top 10 account for 81.6%.
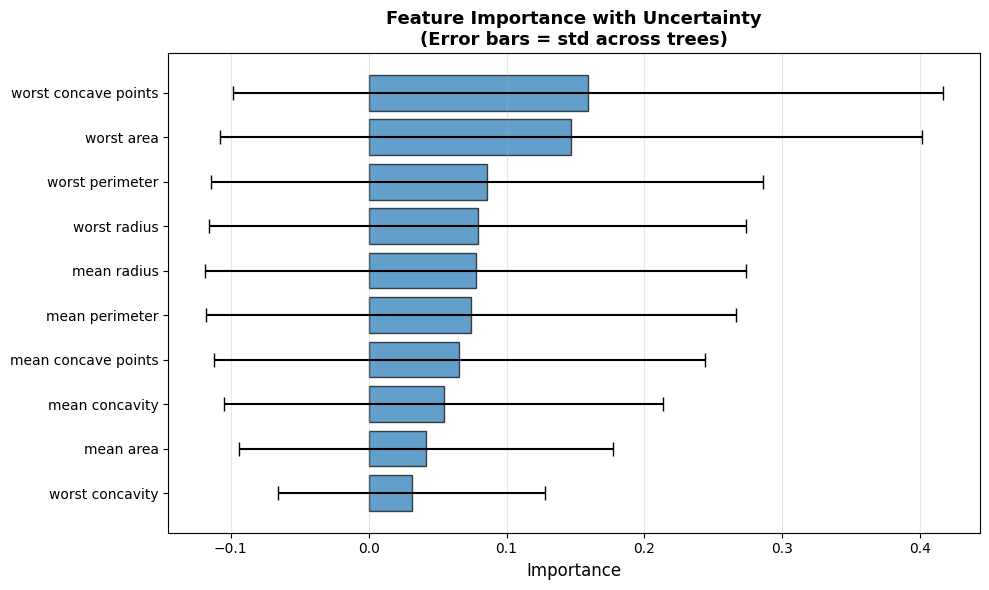
6.4.6.3. Visualizing Feature Importance with Standard Deviation#
tree_importances = np.array([tree.feature_importances_ for tree in rf.estimators_])
importances_mean = tree_importances.mean(axis=0)
importances_std = tree_importances.std(axis=0)
indices = np.argsort(importances_mean)[::-1]
top_10_idx = indices[:10]
plt.figure(figsize=(10, 6))
plt.barh(range(10), importances_mean[top_10_idx], alpha=0.7,
xerr=importances_std[top_10_idx], capsize=5, edgecolor='black')
plt.yticks(range(10), [feature_names[i] for i in top_10_idx], fontsize=10)
plt.xlabel('Importance', fontsize=12)
plt.title('Feature Importance with Uncertainty\n(Error bars = std across trees)',
fontsize=13, fontweight='bold')
plt.gca().invert_yaxis()
plt.grid(True, alpha=0.3, axis='x')
plt.tight_layout()
plt.show()
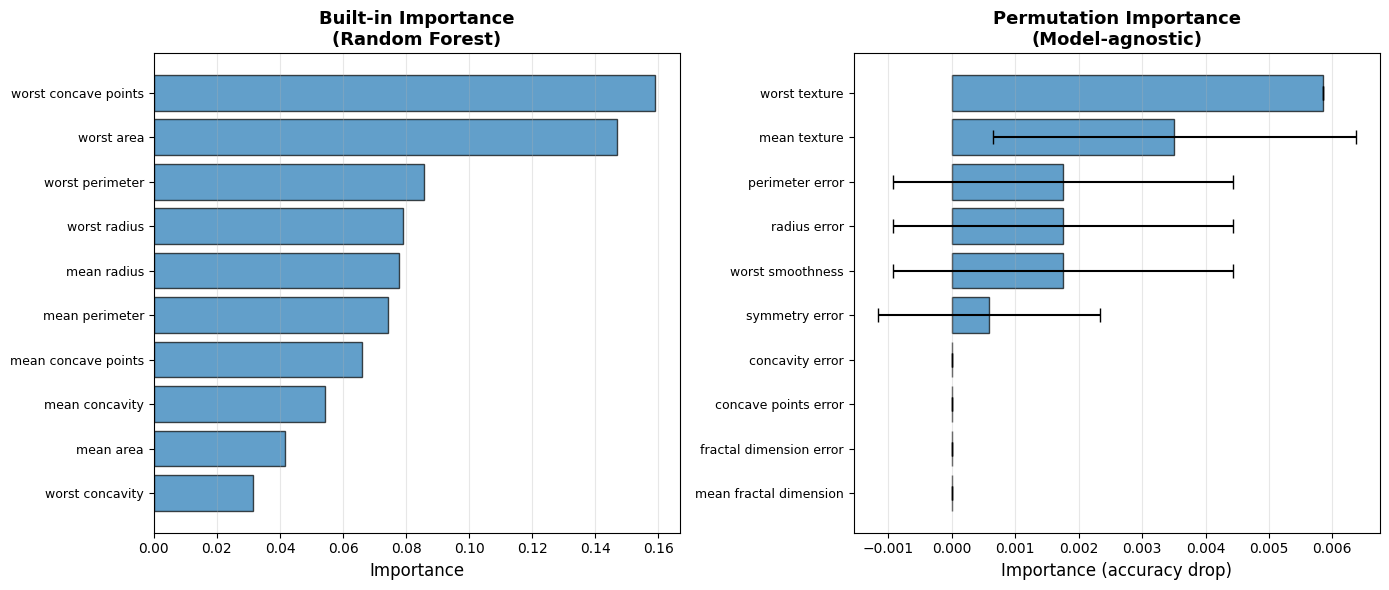
6.4.6.4. Permutation Importance: Model-Agnostic#
Permutation importance: Shuffle feature values, measure performance drop
Algorithm:
Calculate baseline performance
For each feature:
Shuffle feature values (break relationship with target)
Calculate new performance
Importance = baseline - shuffled performance
Repeat multiple times for stability
Pros: Works for any model, accounts for feature interactions Cons: Slower than built-in
perm_importance = permutation_importance(
rf, X_test, y_test, n_repeats=10, random_state=42, n_jobs=-1
)
perm_mean = perm_importance.importances_mean
perm_std = perm_importance.importances_std
perm_indices = np.argsort(perm_mean)[::-1]
perm_top10_df = pd.DataFrame([
{'Rank': i+1, 'Feature': feature_names[perm_indices[i]],
'Mean drop': round(perm_mean[perm_indices[i]], 4),
'Std': round(perm_std[perm_indices[i]], 4)}
for i in range(10)
])
display(perm_top10_df)
fig, axes = plt.subplots(1, 2, figsize=(14, 6))
top_10_builtin = indices[:10]
axes[0].barh(range(10), importances_mean[top_10_builtin], alpha=0.7, edgecolor='black')
axes[0].set_yticks(range(10))
axes[0].set_yticklabels([feature_names[i] for i in top_10_builtin], fontsize=9)
axes[0].set_xlabel('Importance', fontsize=12)
axes[0].set_title('Built-in Importance\n(Random Forest)', fontsize=13, fontweight='bold')
axes[0].invert_yaxis()
axes[0].grid(True, alpha=0.3, axis='x')
top_10_perm = perm_indices[:10]
axes[1].barh(range(10), perm_mean[top_10_perm], alpha=0.7,
xerr=perm_std[top_10_perm], capsize=5, edgecolor='black')
axes[1].set_yticks(range(10))
axes[1].set_yticklabels([feature_names[i] for i in top_10_perm], fontsize=9)
axes[1].set_xlabel('Importance (accuracy drop)', fontsize=12)
axes[1].set_title('Permutation Importance\n(Model-agnostic)', fontsize=13, fontweight='bold')
axes[1].invert_yaxis()
axes[1].grid(True, alpha=0.3, axis='x')
plt.tight_layout()
plt.show()
Traceback (most recent call last):
File "/home/runner/.local/share/uv/python/cpython-3.13.12-linux-x86_64-gnu/lib/python3.13/multiprocessing/resource_tracker.py", line 371, in main
raise ValueError(
f'Cannot register {name} for automatic cleanup: '
f'unknown resource type {rtype}')
ValueError: Cannot register /dev/shm/joblib_memmapping_folder_4407_d8957448d8ab4172a83cd11c9de41fd6_d4b00073e1d74a6b8653fdd57ef85e46 for automatic cleanup: unknown resource type folder
Traceback (most recent call last):
File "/home/runner/.local/share/uv/python/cpython-3.13.12-linux-x86_64-gnu/lib/python3.13/multiprocessing/resource_tracker.py", line 371, in main
raise ValueError(
f'Cannot register {name} for automatic cleanup: '
f'unknown resource type {rtype}')
ValueError: Cannot register /loky-4407-cb8mud1q for automatic cleanup: unknown resource type semlock
Traceback (most recent call last):
File "/home/runner/.local/share/uv/python/cpython-3.13.12-linux-x86_64-gnu/lib/python3.13/multiprocessing/resource_tracker.py", line 371, in main
raise ValueError(
f'Cannot register {name} for automatic cleanup: '
f'unknown resource type {rtype}')
ValueError: Cannot register /loky-4407-zjhbh3e_ for automatic cleanup: unknown resource type semlock
Traceback (most recent call last):
File "/home/runner/.local/share/uv/python/cpython-3.13.12-linux-x86_64-gnu/lib/python3.13/multiprocessing/resource_tracker.py", line 371, in main
raise ValueError(
f'Cannot register {name} for automatic cleanup: '
f'unknown resource type {rtype}')
ValueError: Cannot register /loky-4407-zkckb6ln for automatic cleanup: unknown resource type semlock
Traceback (most recent call last):
File "/home/runner/.local/share/uv/python/cpython-3.13.12-linux-x86_64-gnu/lib/python3.13/multiprocessing/resource_tracker.py", line 371, in main
raise ValueError(
f'Cannot register {name} for automatic cleanup: '
f'unknown resource type {rtype}')
ValueError: Cannot register /loky-4407-2yjks112 for automatic cleanup: unknown resource type semlock
Traceback (most recent call last):
File "/home/runner/.local/share/uv/python/cpython-3.13.12-linux-x86_64-gnu/lib/python3.13/multiprocessing/resource_tracker.py", line 371, in main
raise ValueError(
f'Cannot register {name} for automatic cleanup: '
f'unknown resource type {rtype}')
ValueError: Cannot register /loky-4407-q7ojvifg for automatic cleanup: unknown resource type semlock
Traceback (most recent call last):
File "/home/runner/.local/share/uv/python/cpython-3.13.12-linux-x86_64-gnu/lib/python3.13/multiprocessing/resource_tracker.py", line 371, in main
raise ValueError(
f'Cannot register {name} for automatic cleanup: '
f'unknown resource type {rtype}')
ValueError: Cannot register /loky-4407-iyq19j3c for automatic cleanup: unknown resource type semlock
Traceback (most recent call last):
File "/home/runner/.local/share/uv/python/cpython-3.13.12-linux-x86_64-gnu/lib/python3.13/multiprocessing/resource_tracker.py", line 371, in main
raise ValueError(
f'Cannot register {name} for automatic cleanup: '
f'unknown resource type {rtype}')
ValueError: Cannot register /dev/shm/joblib_memmapping_folder_4407_d8957448d8ab4172a83cd11c9de41fd6_3e853ee70c9540bfb8d16abe76cbda1d for automatic cleanup: unknown resource type folder
Traceback (most recent call last):
File "/home/runner/.local/share/uv/python/cpython-3.13.12-linux-x86_64-gnu/lib/python3.13/multiprocessing/resource_tracker.py", line 371, in main
raise ValueError(
f'Cannot register {name} for automatic cleanup: '
f'unknown resource type {rtype}')
ValueError: Cannot register /loky-4407-_k98az0s for automatic cleanup: unknown resource type semlock
Traceback (most recent call last):
File "/home/runner/.local/share/uv/python/cpython-3.13.12-linux-x86_64-gnu/lib/python3.13/multiprocessing/resource_tracker.py", line 371, in main
raise ValueError(
f'Cannot register {name} for automatic cleanup: '
f'unknown resource type {rtype}')
ValueError: Cannot register /loky-4407-_r67yxe_ for automatic cleanup: unknown resource type semlock
Traceback (most recent call last):
File "/home/runner/.local/share/uv/python/cpython-3.13.12-linux-x86_64-gnu/lib/python3.13/multiprocessing/resource_tracker.py", line 371, in main
raise ValueError(
f'Cannot register {name} for automatic cleanup: '
f'unknown resource type {rtype}')
ValueError: Cannot register /dev/shm/joblib_memmapping_folder_4407_d8957448d8ab4172a83cd11c9de41fd6_3e853ee70c9540bfb8d16abe76cbda1d for automatic cleanup: unknown resource type folder
| Rank | Feature | Mean drop | Std | |
|---|---|---|---|---|
| 0 | 1 | worst texture | 0.0058 | 0.0000 |
| 1 | 2 | mean texture | 0.0035 | 0.0029 |
| 2 | 3 | perimeter error | 0.0018 | 0.0027 |
| 3 | 4 | radius error | 0.0018 | 0.0027 |
| 4 | 5 | worst smoothness | 0.0018 | 0.0027 |
| 5 | 6 | symmetry error | 0.0006 | 0.0018 |
| 6 | 7 | concavity error | 0.0000 | 0.0000 |
| 7 | 8 | concave points error | 0.0000 | 0.0000 |
| 8 | 9 | fractal dimension error | 0.0000 | 0.0000 |
| 9 | 10 | mean fractal dimension | 0.0000 | 0.0000 |
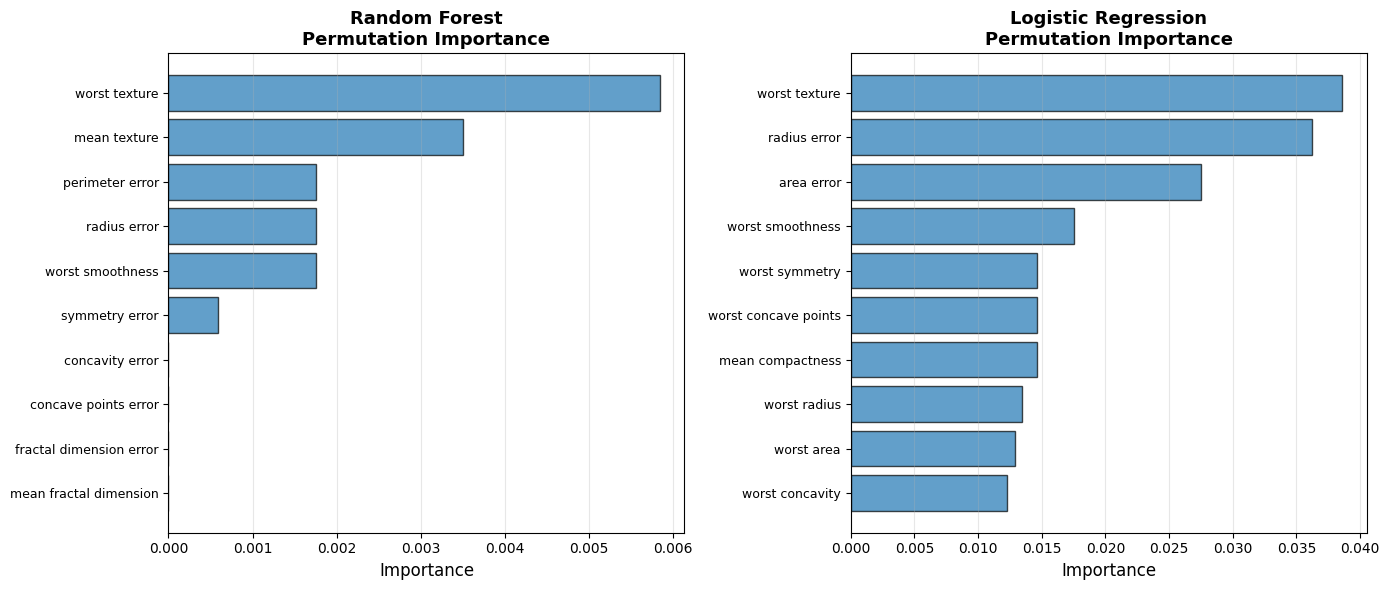
6.4.6.5. Permutation Importance for Any Model#
Key advantage: Works for models without built-in importance!
scaler = StandardScaler()
X_train_scaled = scaler.fit_transform(X_train)
X_test_scaled = scaler.transform(X_test)
lr_model = LogisticRegression(max_iter=10000, random_state=42)
lr_model.fit(X_train_scaled, y_train)
perm_import_lr = permutation_importance(lr_model, X_test_scaled, y_test, n_repeats=10, random_state=42)
perm_mean_lr = perm_import_lr.importances_mean
perm_indices_lr = np.argsort(perm_mean_lr)[::-1]
fig, axes = plt.subplots(1, 2, figsize=(14, 6))
top_10_rf = perm_indices[:10]
axes[0].barh(range(10), perm_mean[top_10_rf], alpha=0.7, edgecolor='black')
axes[0].set_yticks(range(10))
axes[0].set_yticklabels([feature_names[i] for i in top_10_rf], fontsize=9)
axes[0].set_xlabel('Importance', fontsize=12)
axes[0].set_title('Random Forest\nPermutation Importance', fontsize=13, fontweight='bold')
axes[0].invert_yaxis()
axes[0].grid(True, alpha=0.3, axis='x')
top_10_lr = perm_indices_lr[:10]
axes[1].barh(range(10), perm_mean_lr[top_10_lr], alpha=0.7, edgecolor='black')
axes[1].set_yticks(range(10))
axes[1].set_yticklabels([feature_names[i] for i in top_10_lr], fontsize=9)
axes[1].set_xlabel('Importance', fontsize=12)
axes[1].set_title('Logistic Regression\nPermutation Importance', fontsize=13, fontweight='bold')
axes[1].invert_yaxis()
axes[1].grid(True, alpha=0.3, axis='x')
plt.tight_layout()
plt.show()
6.4.6.6. Regression Example: Feature Importance#
from sklearn.ensemble import RandomForestRegressor
diabetes = load_diabetes()
X_diab, y_diab = diabetes.data, diabetes.target
feature_names_diab = diabetes.feature_names
X_train_diab, X_test_diab, y_train_diab, y_test_diab = train_test_split(
X_diab, y_diab, test_size=0.3, random_state=42
)
rf_reg = RandomForestRegressor(n_estimators=100, random_state=42)
rf_reg.fit(X_train_diab, y_train_diab)
importances_diab = rf_reg.feature_importances_
indices_diab = np.argsort(importances_diab)[::-1]
display(pd.DataFrame([
{'Rank': i+1, 'Feature': feature_names_diab[indices_diab[i]],
'Importance': round(importances_diab[indices_diab[i]], 4)}
for i in range(len(feature_names_diab))
]))
plt.figure(figsize=(10, 6))
plt.barh(range(len(feature_names_diab)), importances_diab[indices_diab], alpha=0.7, edgecolor='black')
plt.yticks(range(len(feature_names_diab)), [feature_names_diab[i] for i in indices_diab])
plt.xlabel('Importance', fontsize=12)
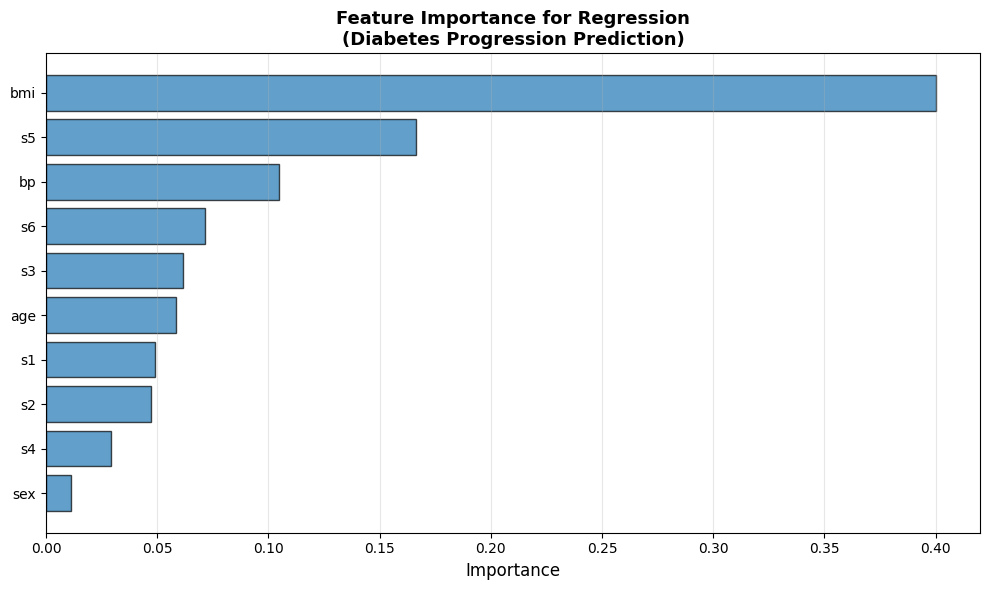
plt.title('Feature Importance for Regression\n(Diabetes Progression Prediction)', fontsize=13, fontweight='bold')
plt.gca().invert_yaxis()
plt.grid(True, alpha=0.3, axis='x')
plt.tight_layout()
plt.show()
| Rank | Feature | Importance | |
|---|---|---|---|
| 0 | 1 | bmi | 0.4000 |
| 1 | 2 | s5 | 0.1666 |
| 2 | 3 | bp | 0.1048 |
| 3 | 4 | s6 | 0.0714 |
| 4 | 5 | s3 | 0.0617 |
| 5 | 6 | age | 0.0586 |
| 6 | 7 | s1 | 0.0492 |
| 7 | 8 | s2 | 0.0471 |
| 8 | 9 | s4 | 0.0294 |
| 9 | 10 | sex | 0.0111 |
6.4.6.7. Using Feature Importance for Feature Selection#
Application: Remove unimportant features to simplify model
Benefits:
Faster training
Less overfitting
Easier interpretation
Lower data collection cost
importances_sorted = np.sort(importances)[::-1]
cumsum_importance = np.cumsum(importances_sorted)
n_features_95 = np.argmax(cumsum_importance >= 0.95) + 1
n_selected = 10
selected_features = indices[:n_selected]
X_train_selected = X_train[:, selected_features]
X_test_selected = X_test[:, selected_features]
rf_selected = RandomForestClassifier(n_estimators=100, random_state=42)
rf_selected.fit(X_train_selected, y_train)
acc_all = rf.score(X_test, y_test)
acc_selected = rf_selected.score(X_test_selected, y_test)
glue('fs-n95', n_features_95, display=False)
glue('fs-acc-all', round(acc_all, 3), display=False)
glue('fs-acc-sel', round(acc_selected, 3), display=False)
plt.figure(figsize=(10, 6))
plt.plot(range(1, len(cumsum_importance)+1), cumsum_importance, linewidth=2, marker='o', markersize=4)
plt.axhline(0.95, color='red', linestyle='--', linewidth=2, label='95% threshold')
plt.axvline(n_features_95, color='red', linestyle='--', linewidth=2, label=f'{n_features_95} features needed')
plt.fill_between(range(1, n_features_95+1), cumsum_importance[:n_features_95], alpha=0.3)
plt.xlabel('Number of Features (sorted by importance)', fontsize=12)
plt.ylabel('Cumulative Importance', fontsize=12)
plt.title('Cumulative Feature Importance\nFew features capture most information', fontsize=13, fontweight='bold')
plt.legend()
plt.grid(True, alpha=0.3)
plt.tight_layout()
plt.show()
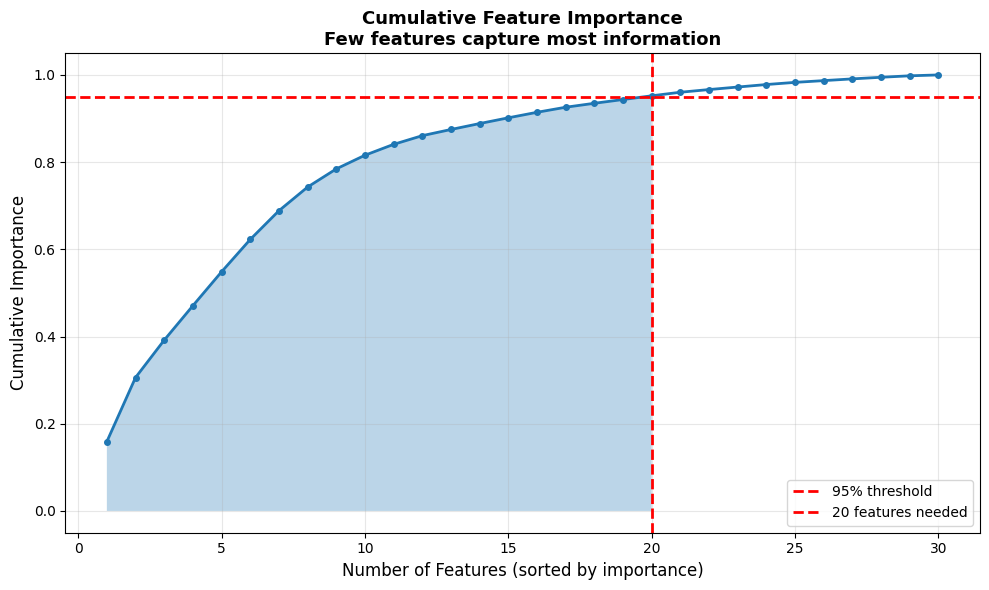
np.int64(20) features reach 95% cumulative importance. Using only the top {n_selected} features: all-features accuracy = 0.936, selected accuracy = 0.942.
6.4.6.8. Detecting Data Leakage with Feature Importance#
Data leakage: Information from target “leaks” into features
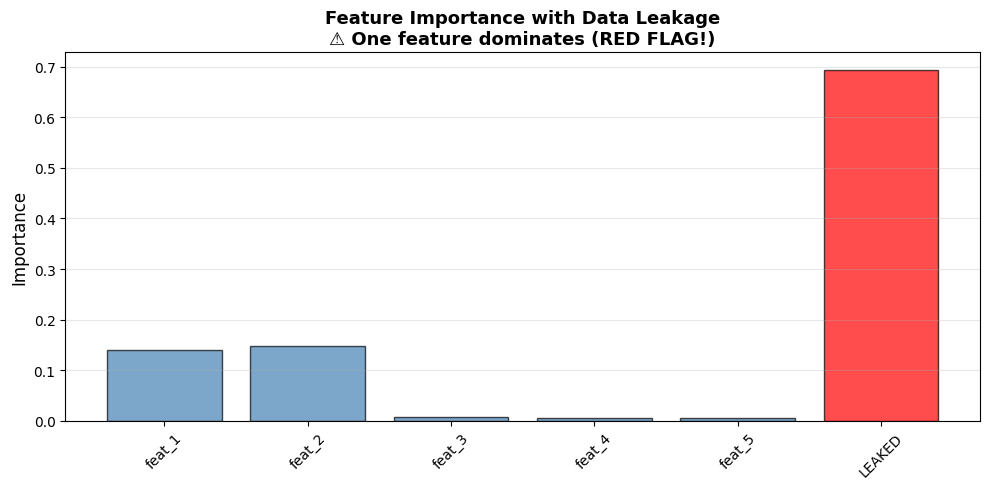
Red flag: Feature suspiciously important (>50% importance)
Example: Using “purchase_date” to predict “will_purchase”
np.random.seed(42)
n_samples = 1000
X_normal = np.random.randn(n_samples, 5)
y_leaked = (X_normal[:, 0] + X_normal[:, 1] > 0).astype(int)
X_leaked_feat = y_leaked + np.random.randn(n_samples) * 0.1
X_with_leak = np.column_stack([X_normal, X_leaked_feat])
feature_names_leak = ['feat_1', 'feat_2', 'feat_3', 'feat_4', 'feat_5', 'LEAKED']
X_train_leak, X_test_leak, y_train_leak, y_test_leak = train_test_split(
X_with_leak, y_leaked, test_size=0.3, random_state=42
)
rf_leak = RandomForestClassifier(n_estimators=100, random_state=42)
rf_leak.fit(X_train_leak, y_train_leak)
importances_leak = rf_leak.feature_importances_
display(pd.DataFrame([
{'Feature': name, 'Importance': round(imp, 4),
'Flag': '⚠️ LEAKED' if name == 'LEAKED' else ''}
for name, imp in zip(feature_names_leak, importances_leak)
]))
colors = ['steelblue'] * 5 + ['red']
plt.figure(figsize=(10, 5))
plt.bar(range(len(importances_leak)), importances_leak, color=colors, alpha=0.7, edgecolor='black')
plt.xticks(range(len(importances_leak)), feature_names_leak, rotation=45)
plt.ylabel('Importance', fontsize=12)
plt.title('Feature Importance with Data Leakage\n⚠️ One feature dominates (RED FLAG!)', fontsize=13, fontweight='bold')
plt.grid(True, alpha=0.3, axis='y')
plt.tight_layout()
plt.show()
| Feature | Importance | Flag | |
|---|---|---|---|
| 0 | feat_1 | 0.1397 | |
| 1 | feat_2 | 0.1481 | |
| 2 | feat_3 | 0.0075 | |
| 3 | feat_4 | 0.0055 | |
| 4 | feat_5 | 0.0051 | |
| 5 | LEAKED | 0.6940 | ⚠️ LEAKED |
6.4.6.9. Key Takeaways#
Important
Remember These Points:
Built-in Importance (Trees)
Fast, automatic
Based on impurity reduction
Only for tree-based models
Permutation Importance
Model-agnostic (works for any model)
More reliable than built-in
Based on performance drop
Feature Selection
Top features capture most information
Can remove 50-90% features safely
Simpler, faster, more robust
Data Leakage Detection
One feature >50% = red flag
Check domain sensibility
Available at prediction time?
Interpretation
High importance ≠ causation
Different models, different importances
Validate with domain experts
Best Practices
Use multiple methods
Check for leakage
Simplify with feature selection
Document findings

