6.1.4. Model Complexity#
In machine learning, one of the most important concepts is model complexity - how flexible or expressive a model can be. Understanding complexity is crucial because it directly determines whether your model will succeed or fail. In this section, we’ll build intuition for what complexity means and why it matters.
6.1.4.1. What is Model Complexity?#
Imagine you’re trying to draw a curve through some points:
Simple approach: Draw a straight line
Complex approach: Draw a wiggly curve that touches every point
Which is better? It depends! This is the essence of model complexity.
Model complexity refers to a model’s capacity to fit various patterns:
Simple models: Limited flexibility, strong assumptions
Complex models: High flexibility, few assumptions
import numpy as np
import matplotlib.pyplot as plt
from sklearn.linear_model import LinearRegression
from sklearn.preprocessing import PolynomialFeatures
from sklearn.pipeline import make_pipeline
from myst_nb import glue
# Generate data with some noise
np.random.seed(42)
X = np.sort(np.random.uniform(0, 10, 30)).reshape(-1, 1)
y = 2 + 3*X.ravel() - 0.5*X.ravel()**2 + np.random.normal(0, 5, 30)
# Create models of different complexity
models = {
'Simple (Linear)': make_pipeline(PolynomialFeatures(1), LinearRegression()),
'Medium (Quadratic)': make_pipeline(PolynomialFeatures(2), LinearRegression()),
'Complex (Degree 9)': make_pipeline(PolynomialFeatures(9), LinearRegression())
}
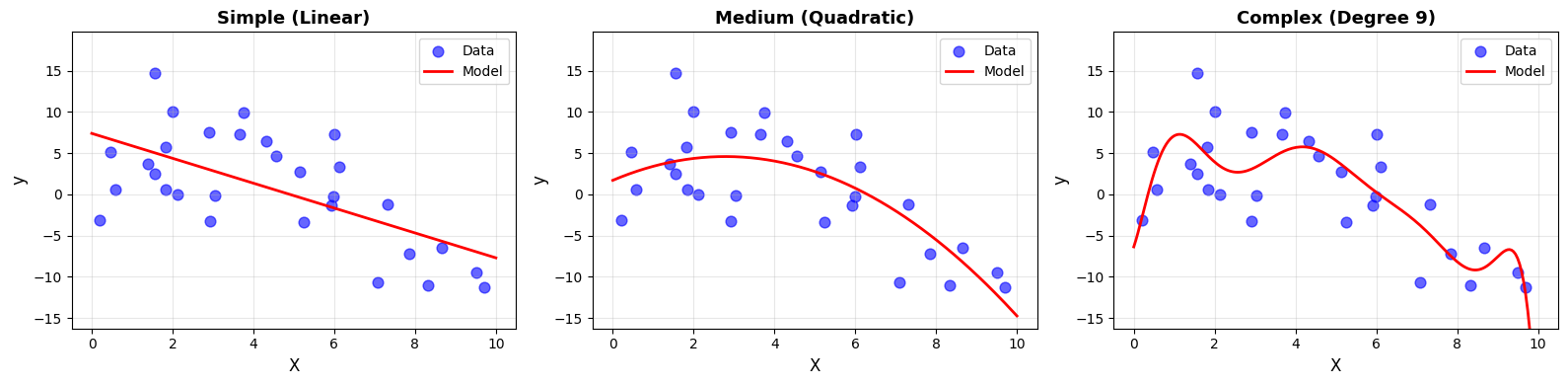
Notice how the complexity affects the fit:
Simple: Smooth but misses curvature
Medium: Captures underlying pattern
Complex: Wiggly, follows noise
6.1.4.2. Defining Complexity#
Model complexity describes how expressive a model is. A more complex model can represent richer patterns, but it also carries a higher risk of overfitting.
There are three closely related ways to understand complexity.
Number of Parameters#
The simplest measure of complexity is the number of learnable parameters.
More parameters → more expressive power
from sklearn.tree import DecisionTreeRegressor
from sklearn.neural_network import MLPRegressor
models_params = [
("Linear (y = mx + b)", LinearRegression(), X, "2 parameters"),
("Degree-5 Polynomial", make_pipeline(PolynomialFeatures(5), LinearRegression()), X, "6 parameters"),
("Decision Tree (depth=10)", DecisionTreeRegressor(max_depth=10, random_state=42), X, "Up to 1024 leaf nodes"),
("Neural Net (10x10)", MLPRegressor(hidden_layer_sizes=(10,10), max_iter=1000, random_state=42), X.ravel().reshape(-1, 1), "~120 parameters")
]
Model Complexity: Number of Parameters
Model |
Parameters |
|---|---|
Linear (y = mx + b) |
2 |
Degree-5 Polynomial |
6 |
Decision Tree (depth=10) |
Up to 1024 leaf nodes |
Neural Net (10x10) |
~120 |
As the number of parameters increases:
The model can capture more detailed patterns
The model becomes more sensitive to noise
The risk of overfitting increases
However, parameter count alone does not fully define complexity.
Model Flexibility#
Flexibility refers to how much the model can bend to follow the data.
Below, polynomial degree controls flexibility.
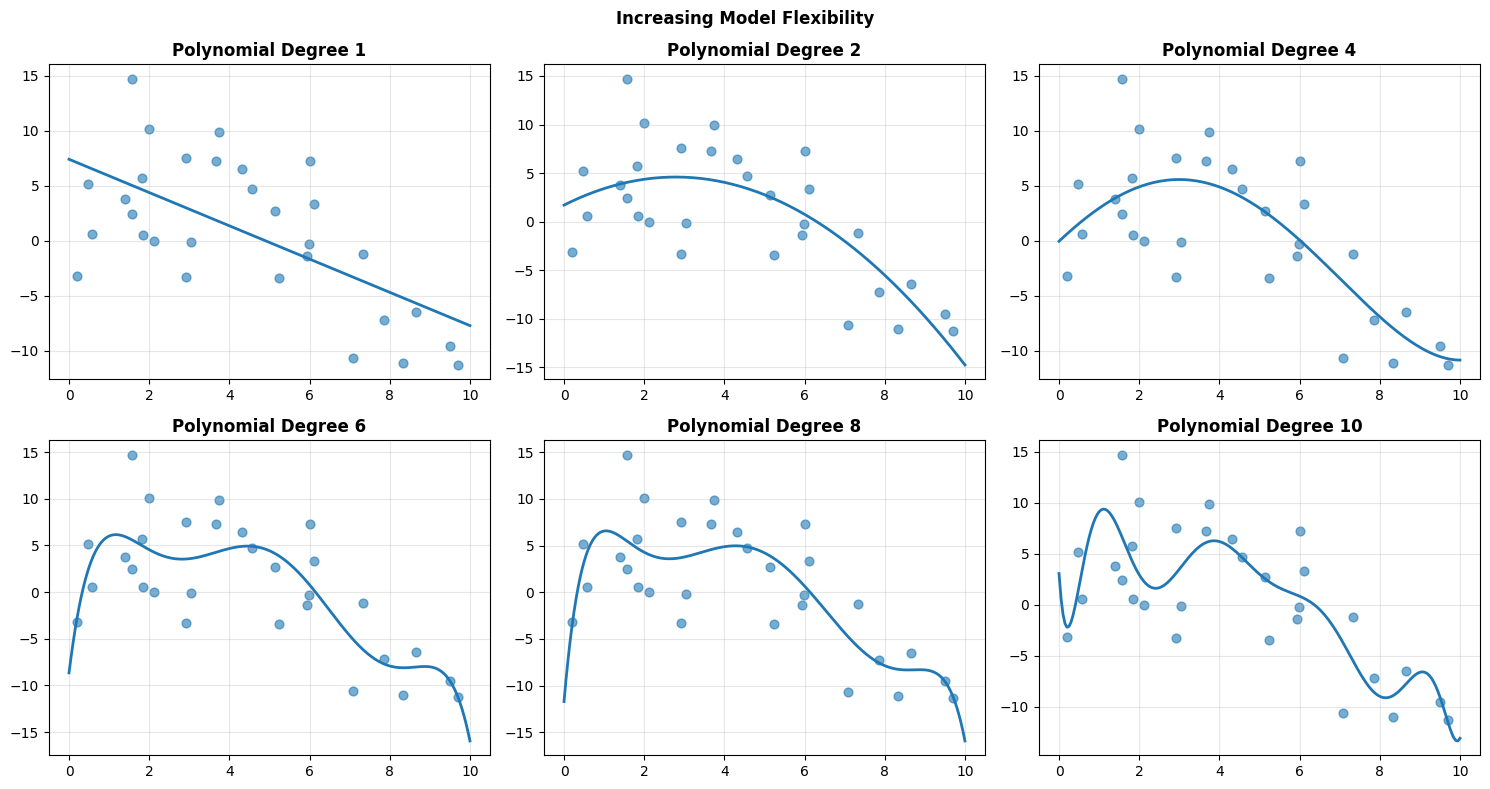
As degree increases:
The curve bends more easily
Training error typically decreases
Sensitivity to noise increases
Low flexibility leads to underfitting. Excessive flexibility leads to overfitting.
Model Capacity#
Capacity refers to the range of functions a model can represent, regardless of whether it actually learns them.
Think of capacity as the model’s vocabulary of patterns.
Low capacity → can express only simple relationships
Medium capacity → can capture moderate curvature
High capacity → can represent highly complex patterns
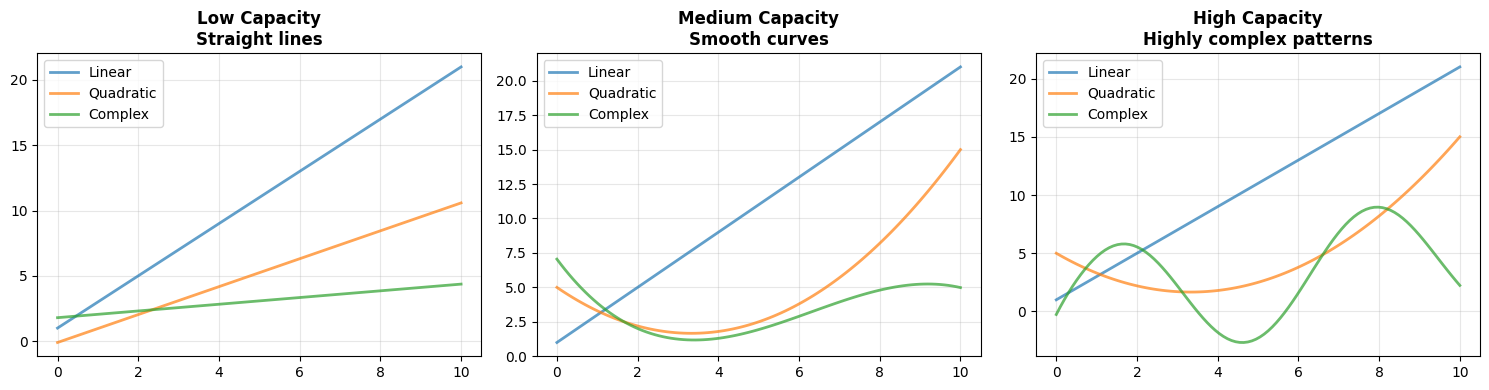
Capacity determines what a model can represent, not necessarily what it will learn.
Low capacity models struggle with complex structure
High capacity models can approximate almost any function
Very high capacity models can also memorize noise
Increasing complexity increases expressive power, but also increases the risk of overfitting. The goal is not to maximize complexity, but to match it appropriately to the underlying structure of the data.
6.1.4.3. Simple Models#
When we begin modeling, we often start with something deliberately modest. A simple model makes strong assumptions about how the world works. In return, it offers clarity, stability, and interpretability.
A linear regression model is the canonical example. It assumes that the relationship between input and output can be described by a straight line.
from sklearn.linear_model import Ridge
from sklearn.linear_model import LinearRegression
# Generate data
np.random.seed(42)
X_simple = np.random.uniform(0, 10, 50).reshape(-1, 1)
y_simple = 3*X_simple.ravel() + 2 + np.random.normal(0, 2, 50)
# Train simple model
model_simple = LinearRegression()
model_simple.fit(X_simple, y_simple)
LinearRegression()In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook.
On GitHub, the HTML representation is unable to render, please try loading this page with nbviewer.org.
Parameters
| fit_intercept | True | |
| copy_X | True | |
| tol | 1e-06 | |
| n_jobs | None | |
| positive | False |
The fitted model is:
[ y = 2.96 * X + 2.19 ]
It contains only two parameters: a slope and an intercept. Its assumption is explicit: the relationship between (X) and (y) is linear.
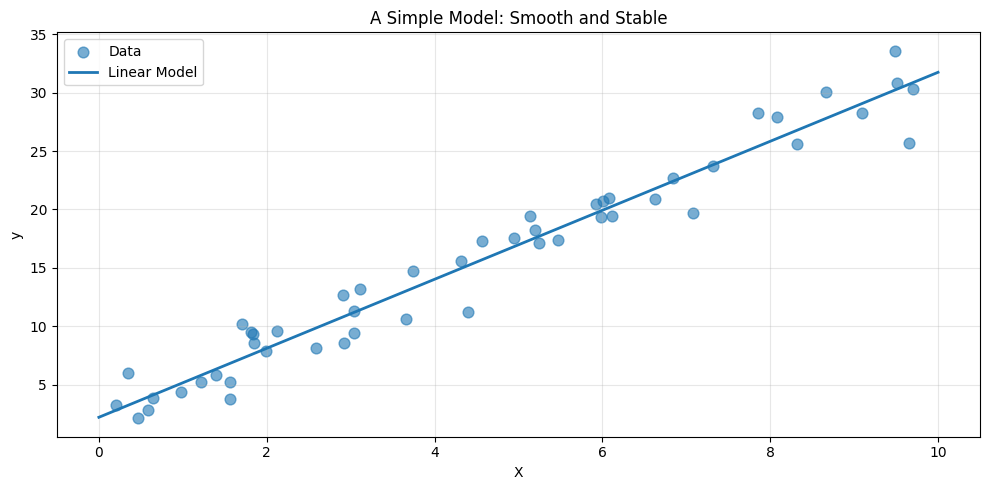
Notice the smoothness of the prediction. The model does not chase individual fluctuations in the data. It captures the overall trend and ignores the noise.
Simple models typically:
Have few parameters
Impose strong structural assumptions
Train quickly
Are easy to interpret
Resist overfitting when data is limited
Their limitation is equally clear. If the true pattern bends or curves, a straight line cannot capture it.
6.1.4.4. Complex Models#
At the other end of the spectrum are flexible, high capacity models. These models make fewer assumptions about structure and allow the data to shape the prediction.
A random forest is one such model.
from sklearn.ensemble import RandomForestRegressor
model_complex = RandomForestRegressor(
n_estimators=100,
max_depth=10,
random_state=42
)
model_complex.fit(X_simple, y_simple)
RandomForestRegressor(max_depth=10, random_state=42)In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook.
On GitHub, the HTML representation is unable to render, please try loading this page with nbviewer.org.
Parameters
| n_estimators | 100 | |
| criterion | 'squared_error' | |
| max_depth | 10 | |
| min_samples_split | 2 | |
| min_samples_leaf | 1 | |
| min_weight_fraction_leaf | 0.0 | |
| max_features | 1.0 | |
| max_leaf_nodes | None | |
| min_impurity_decrease | 0.0 | |
| bootstrap | True | |
| oob_score | False | |
| n_jobs | None | |
| random_state | 42 | |
| verbose | 0 | |
| warm_start | False | |
| ccp_alpha | 0.0 | |
| max_samples | None | |
| monotonic_cst | None |
A random forest is an ensemble of decision trees. Each tree partitions the input space into regions and assigns predictions within those regions. With many trees combined, the model becomes highly expressive.
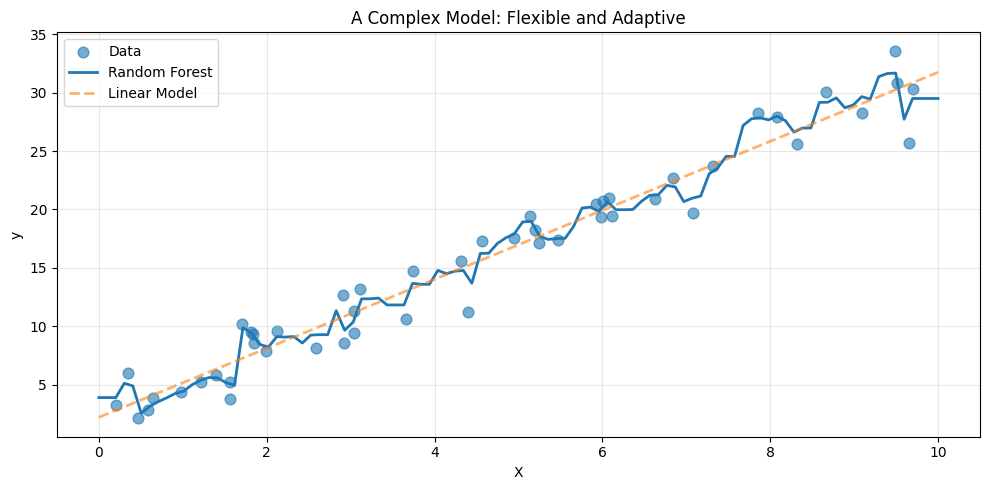
Compared to the linear model, the forest adapts more closely to local variations. It can bend, flatten, and shift depending on the data in each region.
Complex models generally:
Contain many effective parameters
Make weaker assumptions
Capture nonlinear structure
Require more computation
Are harder to interpret
They are powerful, but power comes with risk. With limited data, flexibility can turn into overfitting.
6.1.4.5. The Complexity Spectrum#
Models do not fall into neat categories. They lie along a continuum, from rigid to highly expressive.
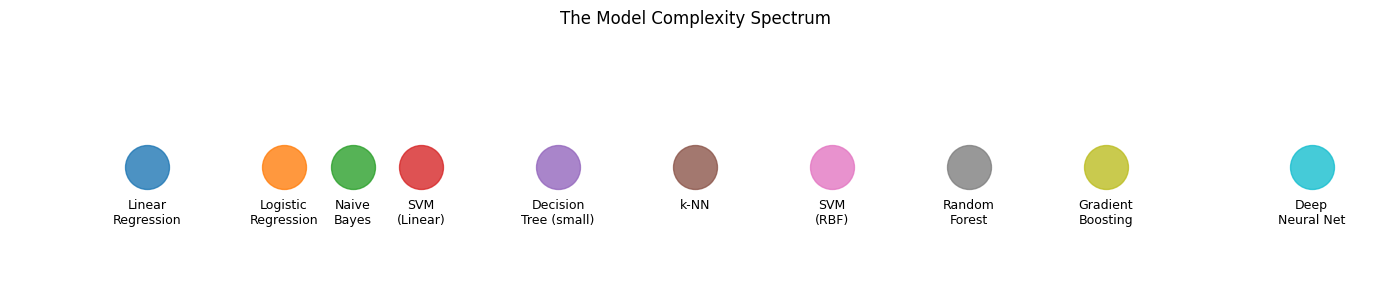

Where a model should sit on this spectrum depends on:
The amount of data available
The true complexity of the underlying pattern
The need for interpretability
Computational constraints
There is no universally best level of complexity. There is only an appropriate match between model and problem.
6.1.4.6. The Goldilocks Principle#
To understand this trade off, consider a dataset generated from a quadratic relationship.
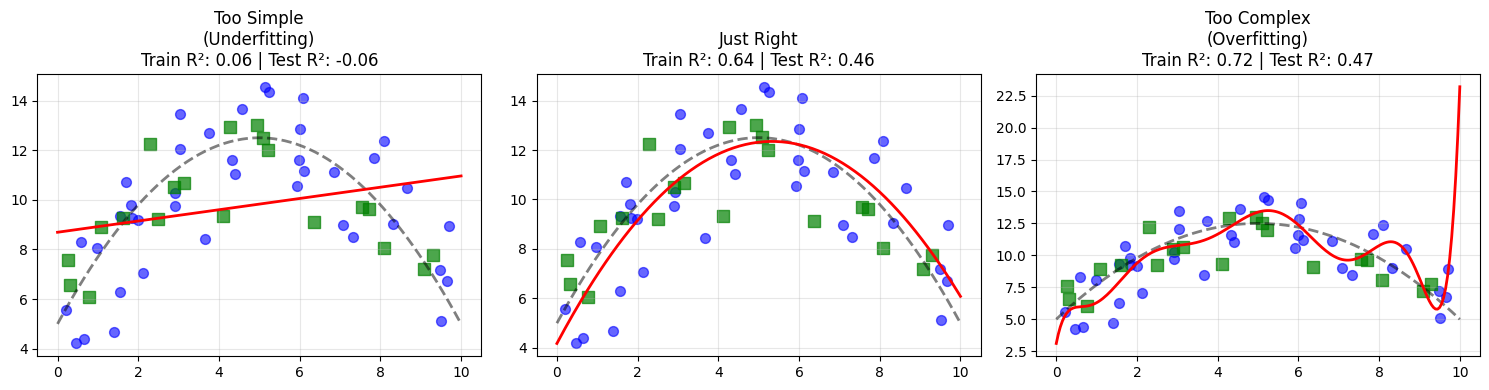
Three regimes emerge:
A degree 1 model underfits. It misses the curvature entirely.
A degree 2 model captures the true structure.
A high degree model fits the noise, achieving near perfect training performance but worse test performance.
This is the Goldilocks principle of modeling. Too simple misses structure. Too complex memorizes noise. The goal is a model that is just complex enough to capture the true signal.
6.1.4.7. Complexity and Performance#
If we increase complexity systematically, a pattern appears.
from sklearn.model_selection import cross_val_score
degrees = range(1, 16)
train_scores = []
test_scores = []
for degree in degrees:
model = make_pipeline(PolynomialFeatures(degree), LinearRegression())
model.fit(X_gold, y_gold)
train_scores.append(model.score(X_gold, y_gold))
test_scores.append(model.score(X_test, y_test))
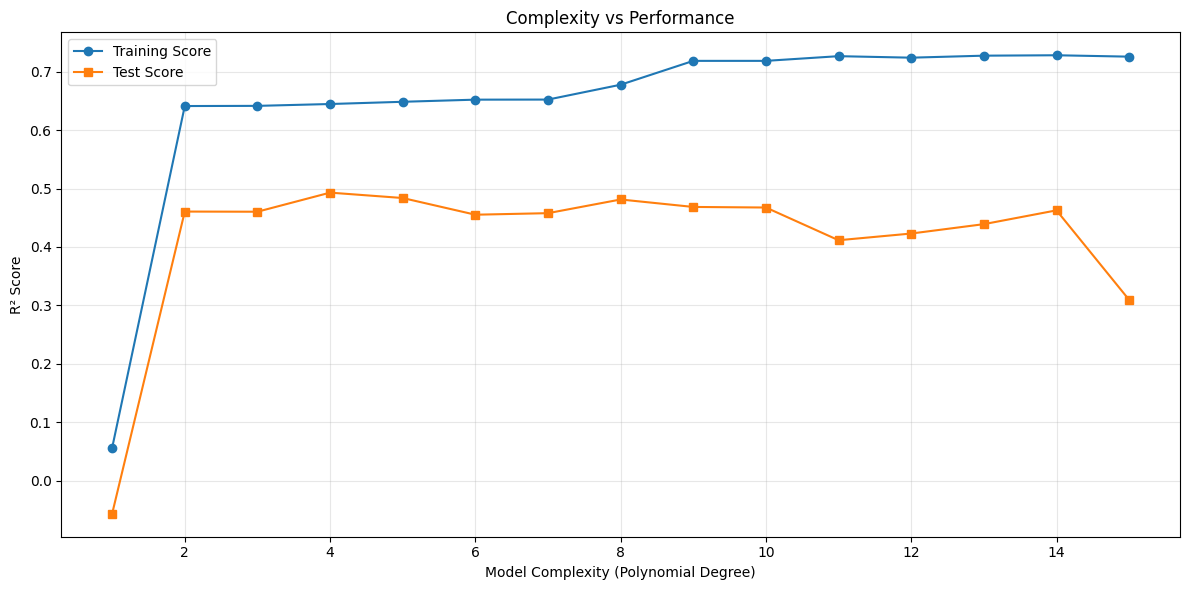
Two consistent observations arise:
Training performance improves monotonically with complexity.
Test performance improves up to a point, then declines.
The optimal model is located at the peak of the test curve. Beyond that point, additional flexibility improves the fit to training data but harms generalization.
6.1.4.8. Choosing the Right Complexity#
In practice:
Begin with the simplest reasonable model.
Increase complexity only when evidence demands it.
Use validation data to monitor generalization.
Watch the gap between training and test performance.
Remember that more data can justify more complex models.
Modeling is not about maximizing flexibility. It is about aligning model capacity with the true structure of the problem.

