6.2.2.7. Random Forest#
Random Forest is the Bagging algorithm applied to classification trees - with one crucial twist: feature randomisation. Instead of just training each tree on a bootstrap sample, at every split the algorithm considers only a random subset of \(m\) features. This decorrelates the trees, making their errors more independent, so the majority vote is far more reliable than any individual tree.
The result is exactly the classification counterpart of Random Forest:
Reduced variance from averaging many diverse trees
Near-zero overfitting compared to a single deep tree
Free out-of-bag (OOB) validation - each tree is tested on the ~37% of data it did not see during training
Random Forest is widely regarded as the best default classifier for tabular data.
The Math#
Training:
For \(b = 1, \ldots, B\):
Draw a bootstrap sample \(\mathcal{D}_b\) (same size as the training set, with replacement).
Grow a full decision tree on \(\mathcal{D}_b\). At every split, sample \(m \leq p\) features and choose the best split only among those \(m\) features.
Store all \(B\) trees.
Prediction (majority vote):
The default choice \(m = \lfloor\sqrt{p}\rfloor\) balances bias and variance in practice.
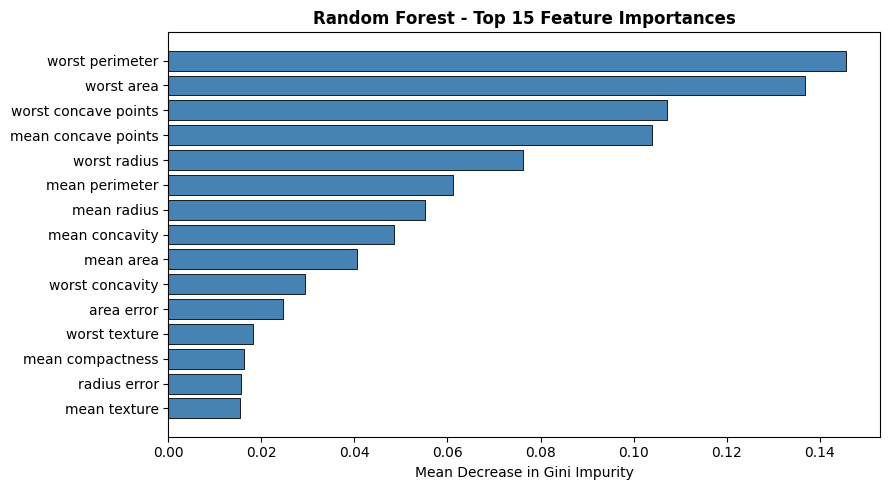
Feature importance is the mean decrease in Gini impurity across all splits on that feature, averaged over all trees.
Key hyperparameters:
Hyperparameter |
Guidance |
|---|---|
|
More trees → better, diminishing returns after ~200-500 |
|
|
|
Default |
|
Smooths individual trees; useful for noisy data |
|
Free out-of-bag validation estimate |
In scikit-learn#
from sklearn.ensemble import RandomForestClassifier
rf = RandomForestClassifier(
n_estimators=200,
max_features='sqrt',
oob_score=True,
random_state=42,
n_jobs=-1
)
rf.fit(X_train, y_train)
No feature scaling is required - Random Forest inherits this property from decision trees.
Example#
from sklearn.ensemble import RandomForestClassifier
rf = RandomForestClassifier(n_estimators=200, max_features='sqrt',
oob_score=True, random_state=42, n_jobs=-1)
rf.fit(X_train, y_train)
train_acc = accuracy_score(y_train, rf.predict(X_train))
test_acc = accuracy_score(y_test, rf.predict(X_test))
test_auc = roc_auc_score(y_test, rf.predict_proba(X_test)[:, 1])
Random Forest achieves a test accuracy of 0.958 and AUC-ROC of 0.995. The OOB accuracy of 0.962 closely tracks the held-out test score - a useful sanity check that doesn’t require a separate validation split. Train accuracy of 1.0 is near perfect (full-depth trees memorise their bootstrap samples) but that does not imply the ensemble overfits.
Feature Importance#
imp_df = pd.DataFrame({
"Feature": data.feature_names,
"Importance": rf.feature_importances_,
}).sort_values("Importance", ascending=True).tail(15)
fig, ax = plt.subplots(figsize=(9, 5))
ax.barh(imp_df["Feature"], imp_df["Importance"],
color='steelblue', edgecolor='black', linewidth=0.6)
ax.set_xlabel("Mean Decrease in Gini Impurity")
ax.set_title("Random Forest - Top 15 Feature Importances", fontweight='bold')
plt.tight_layout()
plt.show()
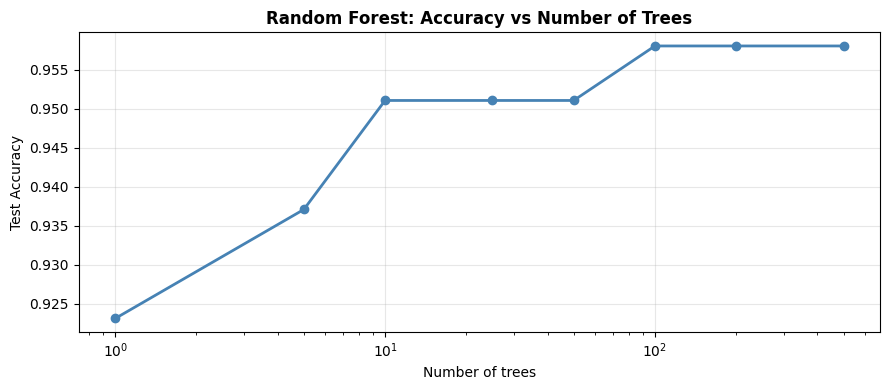
Number of Trees vs Accuracy#
More trees always helps (or at worst, is neutral). Performance plateaus around 100–300 trees on most datasets.
n_values = [1, 5, 10, 25, 50, 100, 200, 500]
test_accs = []
for n in n_values:
m = RandomForestClassifier(n_estimators=n, max_features='sqrt',
random_state=42, n_jobs=-1)
m.fit(X_train, y_train)
test_accs.append(accuracy_score(y_test, m.predict(X_test)))
fig, ax = plt.subplots(figsize=(9, 4))
ax.semilogx(n_values, test_accs, 'o-', color='steelblue', lw=2)
ax.set_xlabel("Number of trees")
ax.set_ylabel("Test Accuracy")
ax.set_title("Random Forest: Accuracy vs Number of Trees", fontweight='bold')
ax.grid(True, alpha=0.3)
plt.tight_layout()
plt.show()