6.2.1.4. Support Vector Regression#
Support Vector Regression (SVR) extends the Support Vector Machine idea from classification to continuous outputs. The key intuition is different from ordinary least squares: instead of minimising the total squared error for every point, SVR tries to fit the data within a tube of width \(\varepsilon\) around the prediction.
Points inside the tube incur zero loss - small errors are tolerated.
Points outside the tube are penalised linearly with distance from the tube boundary.
This tolerance zone makes SVR naturally robust to small amounts of noise. The points that actually sit on or outside the tube boundary are the support vectors - they alone determine the model. All other points are irrelevant to the solution.
The second powerful feature of SVR is the kernel trick: by applying a non-linear transformation to the input space, SVR can fit curved, complex relationships without explicitly engineering polynomial or interaction features.
The Math#
SVR solves the following optimisation problem:
subject to:
where \(\xi_i, \xi_i^* \geq 0\) are slack variables that allow points to sit outside the tube.
The RBF (Radial Basis Function) kernel maps inputs into an infinite-dimensional space, enabling non-linear fits:
Key hyperparameters:
Hyperparameter |
Role |
|---|---|
|
Regularization - high C fits training data tightly; low C is smoother |
|
Tube half-width - errors within this band are ignored entirely |
|
Feature transformation: |
|
RBF width - high gamma → narrow Gaussians, complex boundary |
In scikit-learn#
SVR requires feature scaling - the optimisation is sensitive to feature magnitudes. Always wrap it in a Pipeline with StandardScaler.
from sklearn.svm import SVR
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import StandardScaler
svr = Pipeline([
('scaler', StandardScaler()),
('svr', SVR(kernel='rbf', C=100, epsilon=5))
])
svr.fit(X_train, y_train)
Example#
svr = Pipeline([
('scaler', StandardScaler()),
('svr', SVR(kernel='rbf', C=100, epsilon=5))
])
svr.fit(X_train, y_train)
train_r2 = r2_score(y_train, svr.predict(X_train))
test_r2 = r2_score(y_test, svr.predict(X_test))
test_rmse = np.sqrt(mean_squared_error(y_test, svr.predict(X_test)))
print(f"Train R² : {train_r2:.3f}")
print(f"Test R² : {test_r2:.3f}")
print(f"Test RMSE: {test_rmse:.1f}")
Train R² : 0.945
Test R² : 0.853
Test RMSE: 68.1
SVR achieves a test \(R^2\) of 0.853 and RMSE of 68.1. The train \(R^2\) is 0.945.
The \(\varepsilon\)-Tube on 1-D Data#
To understand how the tube and support vectors work, it is easiest to visualise SVR on a one-dimensional sine curve:
np.random.seed(1)
X_1d = np.sort(np.random.uniform(0, 6, 60)).reshape(-1, 1)
y_1d = 2 * np.sin(X_1d.ravel()) + np.random.normal(0, 0.4, 60)
svr_demo = SVR(kernel='rbf', C=10, epsilon=0.3)
svr_demo.fit(X_1d, y_1d)
Xp = np.linspace(0, 6, 300).reshape(-1, 1)
yp = svr_demo.predict(Xp)
plt.figure(figsize=(10, 5))
plt.scatter(X_1d, y_1d, zorder=3, edgecolors='k', linewidths=0.5,
alpha=0.7, label='Training data', color='steelblue')
plt.plot(Xp, yp, 'r-', linewidth=2.5, label='SVR prediction', zorder=4)
plt.fill_between(Xp.ravel(), yp - 0.3, yp + 0.3,
alpha=0.2, color='red', label='ε-tube (ε=0.3)')
sv_mask = np.zeros(len(X_1d), dtype=bool)
sv_mask[svr_demo.support_] = True
plt.scatter(X_1d[sv_mask], y_1d[sv_mask], s=120, facecolors='none',
edgecolors='blue', linewidths=2, zorder=5,
label=f'Support vectors ({sv_mask.sum()})')
plt.xlabel('X', fontsize=12)
plt.ylabel('y', fontsize=12)
plt.title('SVR - ε-Tube and Support Vectors', fontsize=13, fontweight='bold')
plt.legend(fontsize=10)
plt.grid(True, alpha=0.3)
plt.tight_layout()
plt.show()
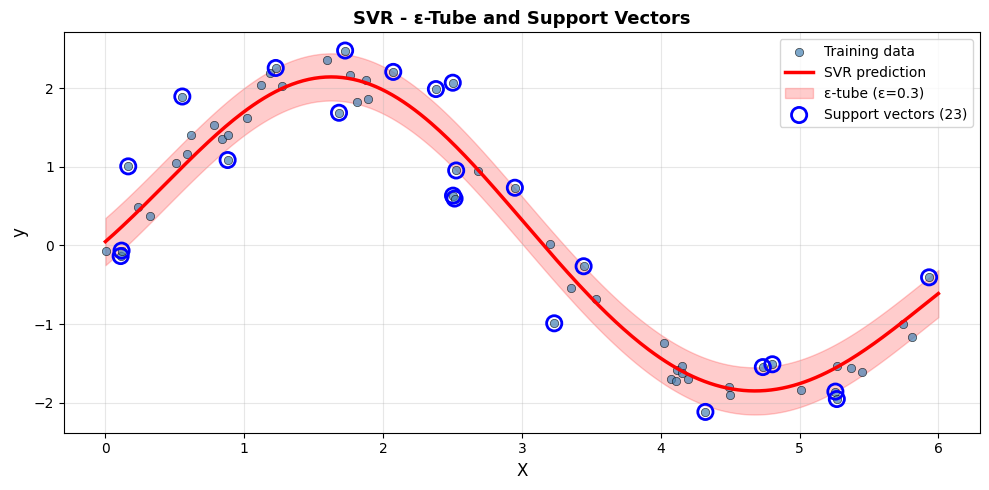
Only 23 of the 60 training points (38.3%) become support vectors. The rest lie inside the tube and have no effect on the model - this is the “support” in SVR.
Hyperparameter Sensitivity - C and Epsilon#
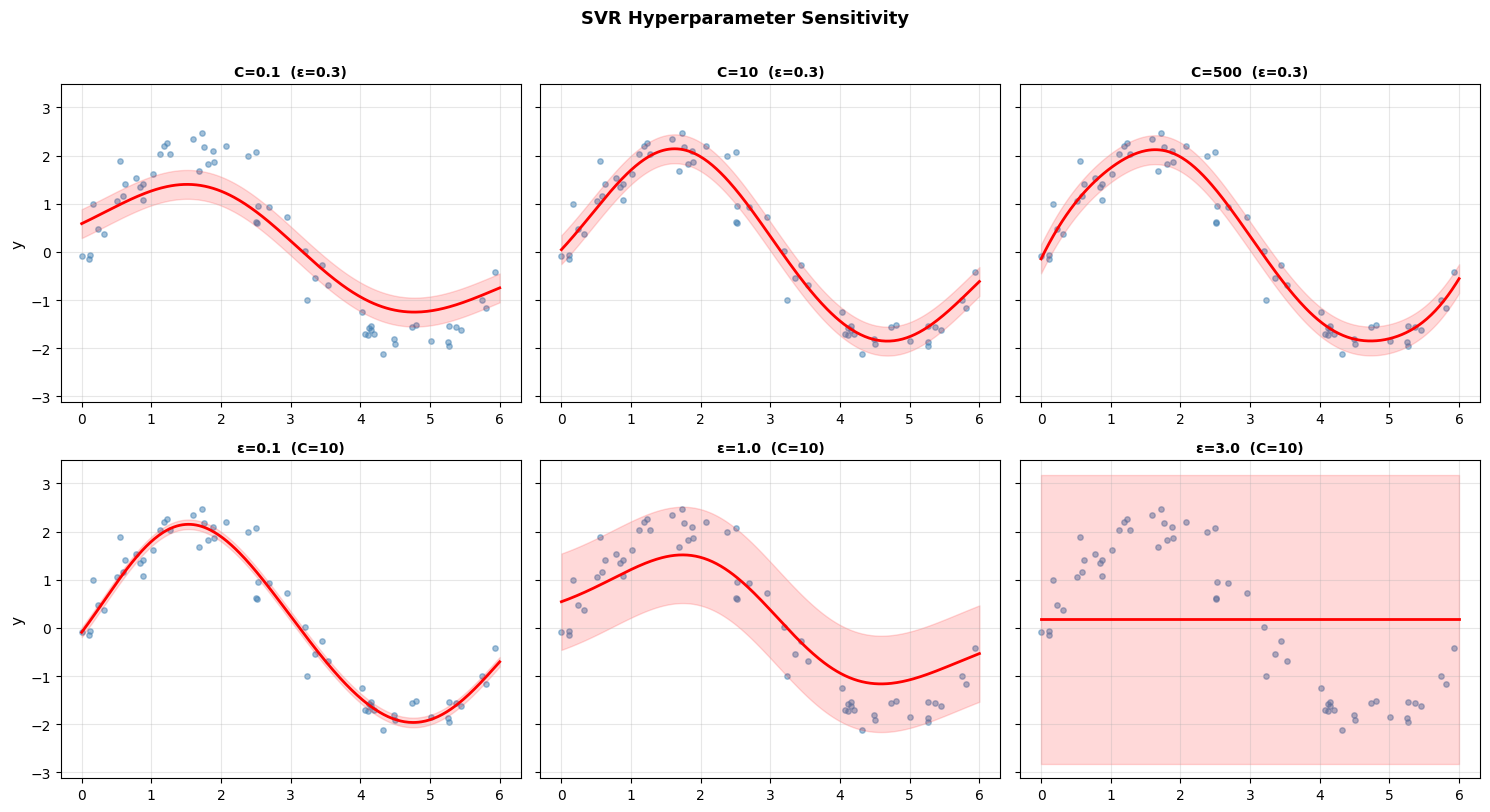
Top row (varying C): Low C produces a smooth, underfit curve. High C fits the training data very tightly, risking overfitting.
Bottom row (varying ε): Small ε uses many support vectors (narrow tolerance). Large ε uses fewer and produces a smoother, coarser fit.
Strengths and Weaknesses#
Strengths |
Handles non-linear relationships via kernels; robust with the ε-tube; effective in high-dimensional spaces |
Weaknesses |
Slow on large datasets (\(O(n^2)\)–\(O(n^3)\)); requires feature scaling; less interpretable; hyperparameter tuning critical |
Tip
SVR excels on small-to-medium datasets (< 10k samples) with non-linear relationships. For large datasets consider Gradient Boosting or Random Forest instead.

