6.2.1.7. Random Forest#
Random Forest is the definitive Bagging algorithm - and the go-to non-linear model for most tabular regression problems. It starts from the same idea as Bagging (see Ensemble Methods): grow many decision trees on bootstrap samples and average their predictions. But it adds one critical twist that makes it far more powerful: feature randomisation.
At each split inside every tree, the algorithm considers only a random subset of \(m\) features (not all \(p\) features). This decorrelates the trees. Without feature randomisation, if one feature is a very strong predictor, almost every tree would split on it first - the trees would look very similar and averaging them would provide little benefit. By forcing each tree to use a random feature subset, trees must find different paths through the data and end up making more independent errors.
The result is:
Reduced variance from averaging many diverse trees
Near-zero overfitting - unlike a single deep tree, the forest is robust
Free validation via out-of-bag (OOB) samples - each tree is validated on the ~37% of data it did not see during training
The Math#
Training algorithm:
For \(b = 1, \ldots, B\):
Draw a bootstrap sample \(\mathcal{D}_b\) of size \(n\) from the training data.
Grow a full (deep) decision tree on \(\mathcal{D}_b\). At every split, randomly sample \(m \leq p\) features and find the best split only among those \(m\) features.
Store all \(B\) trees.
Prediction:
The default choice of \(m = \sqrt{p}\) balances bias and variance well in practice.
Feature importance is computed as the mean decrease in impurity (MSE) across all splits on that feature, averaged over all trees.
Key hyperparameters:
Hyperparameter |
Guidance |
|---|---|
|
More is better up to a point; 100–500 is usually sufficient |
|
|
|
Default |
|
Higher → smoother individual trees; useful for noisy targets |
|
Enables a free out-of-bag validation estimate |
In scikit-learn#
from sklearn.ensemble import RandomForestRegressor
rf = RandomForestRegressor(
n_estimators=200,
max_features='sqrt',
oob_score=True,
random_state=42,
n_jobs=-1 # use all CPU cores
)
rf.fit(X_train, y_train)
print(f"OOB R²: {rf.oob_score_:.3f}") # free estimate of test performance
No feature scaling is needed - Random Forest inherits this property from decision trees.
Example#
rf = RandomForestRegressor(n_estimators=200, max_features='sqrt',
oob_score=True, random_state=42, n_jobs=-1)
rf.fit(X_train, y_train)
train_r2 = r2_score(y_train, rf.predict(X_train))
test_r2 = r2_score(y_test, rf.predict(X_test))
test_rmse = np.sqrt(mean_squared_error(y_test, rf.predict(X_test)))
oob_r2 = rf.oob_score_
print(f"Train R² : {train_r2:.3f}")
print(f"Test R² : {test_r2:.3f}")
print(f"OOB R² : {oob_r2:.3f} ← free estimate, no test data used")
print(f"Test RMSE: {test_rmse:.1f}")
Train R² : 0.963
Test R² : 0.713
OOB R² : 0.732 ← free estimate, no test data used
Test RMSE: 95.3
The forest achieves a test \(R^2\) of 0.713 and an OOB \(R^2\) of 0.732. Notice how closely OOB and test performance agree - the OOB estimate is a reliable free substitute for a separate validation set.
Feature Importance#
Random Forest provides interpretable feature importance scores as a by-product of training - no extra computation required:
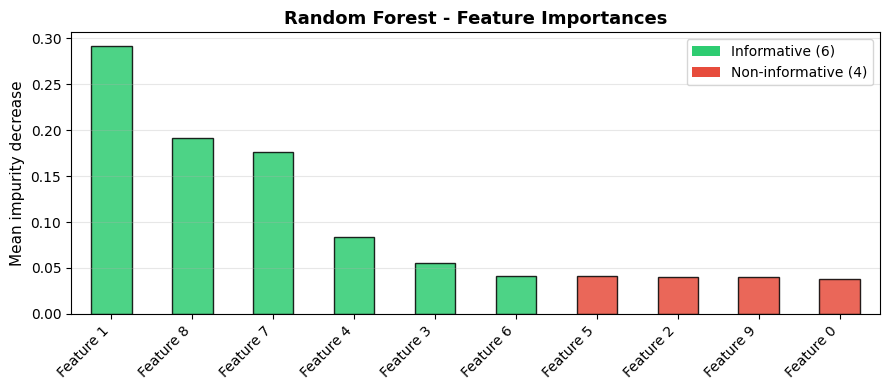
The most important feature is Feature 1 with an importance score of 0.292. The 4 non-informative features (shown in red) cluster at the bottom - Random Forest effectively deprioritises them.
Performance Stabilises with More Trees#
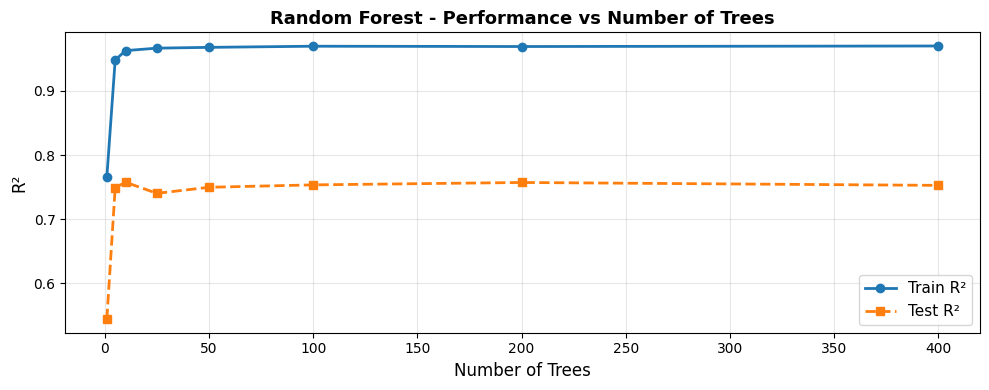
Test performance plateaus around 100–200 trees. Adding more trees beyond this point yields diminishing returns - the forest is already stable.
Effect of max_features on the Bias–Variance Balance#
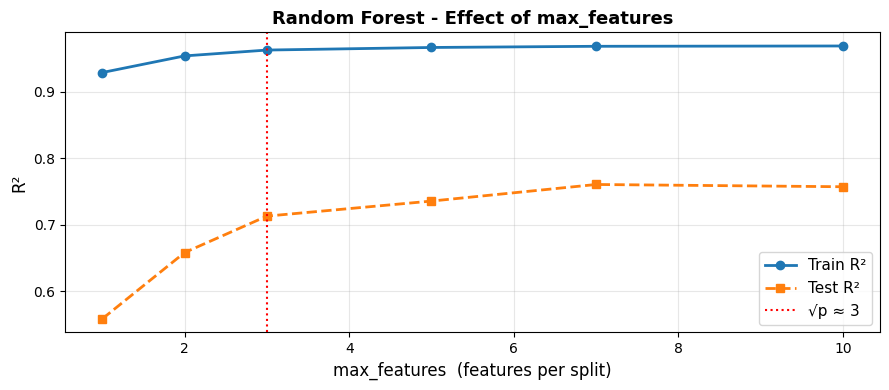
Using fewer features per split (left side) makes trees more diverse but individually weaker - higher bias, lower variance. Using all features (max_features=10) removes the decorrelation benefit. The default \(\sqrt{p}\) sits near the sweet spot.
Strengths and Weaknesses#
Strengths |
Excellent out-of-the-box performance; handles non-linearity; free OOB validation; robust to outliers and missing values; feature importance |
Weaknesses |
Less interpretable than a single tree; slower to train/predict than linear models; can overfit on very noisy data with too many trees |
Tip
Random Forest is one of the best default models for tabular regression. Start here when a linear model under-performs and you need non-linear power with minimal tuning.

