6.1.7. Regularization: Taming Complexity#
In the previous sections, we explored overfitting and the bias–variance tradeoff. We saw how models that are too flexible can memorize noise instead of learning structure. Regularization is the natural next step. It provides a principled way to control complexity and guide models toward better generalization.
You can think of regularization as adding guardrails to a powerful model. A flexible model is capable of fitting almost anything, including randomness. Regularization keeps that flexibility in check without removing it entirely.
6.1.7.1. The Core Idea#
At its heart, regularization introduces a simple but powerful modification to the learning objective:
Important
Regularization = Penalizing Complexity
Instead of just minimizing error, we minimize: \( \text{Total Loss} = \text{Error} + \lambda \times \text{Complexity Penalty}\)
Where:
Error: How well the model fits training data
Complexity Penalty: How complex/flexible the model is
λ (lambda): Controls the tradeoff (hyperparameter)
Rather than asking only, “How well does the model fit the training data?”, we also ask, “How complex is this model?” The hyperparameter λ determines how strongly we care about simplicity. A larger value places more weight on the penalty, encouraging simpler models.
In practice:
Without regularization, the model minimizes only error.
With regularization, the model minimizes error plus a penalty.
The result is often better generalization to unseen data.
The following experiment illustrates this effect concretely.
This forces the model to find a balance between fitting the data and staying simple.
# Generate synthetic data
X = np.sort(np.random.uniform(0, 10, 30)).reshape(-1, 1)
true_function = lambda x: 5 + 2*x - 0.3*x**2
y = true_function(X.ravel()) + np.random.normal(0, 3, 30)
# Split data
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=42)
# Create polynomial features
poly = PolynomialFeatures(10, include_bias=False)
X_train_poly = poly.fit_transform(X_train)
X_test_poly = poly.transform(X_test)
# Train models with and without regularization
model_unreg = LinearRegression()
model_unreg.fit(X_train_poly, y_train)
model_reg = Ridge(alpha=10.0)
model_reg.fit(X_train_poly, y_train)
/home/runner/work/datasciencethenovel/datasciencethenovel/.venv/lib/python3.13/site-packages/scipy/_lib/_util.py:1233: LinAlgWarning: Ill-conditioned matrix (rcond=1.1098e-19): result may not be accurate.
return f(*arrays, *other_args, **kwargs)
Ridge(alpha=10.0)In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook.
On GitHub, the HTML representation is unable to render, please try loading this page with nbviewer.org.
Parameters
| alpha | 10.0 | |
| fit_intercept | True | |
| copy_X | True | |
| max_iter | None | |
| tol | 0.0001 | |
| solver | 'auto' | |
| positive | False | |
| random_state | None |
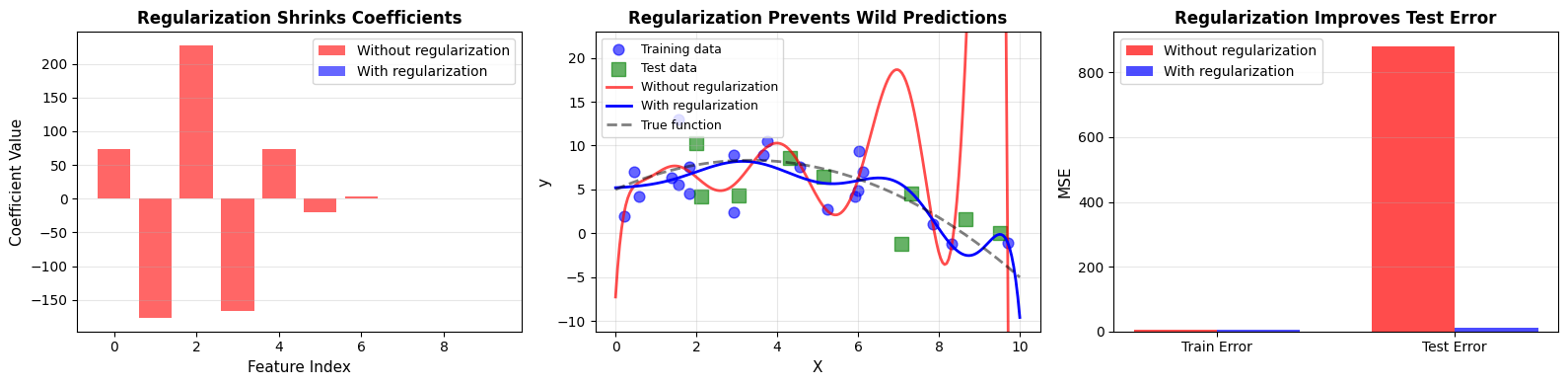
After training both models, we observe a clear pattern:
Without regularization:
Max coefficient magnitude: 226.94
Train MSE: 3.84
Test MSE: 880.95
With regularization:
Max coefficient magnitude: 0.22
Train MSE: 6.35
Test MSE: 11.12
Regularization reduces coefficient magnitudes and improves test performance. The model becomes less extreme, and as a result, more reliable.
6.1.7.2. Why Regularization Works#
Overfitting arises when a model is too flexible relative to the amount of available data. High flexibility means many parameters. With limited data, those parameters can take extreme values to perfectly match noise.
Regularization counteracts this tendency.
The mechanism is intuitive:
Complex models contain many parameters.
With limited data, parameters may grow very large to fit small fluctuations.
Regularization penalizes large parameter values.
The model still captures structure but avoids chasing noise.
The penalty acts as a soft constraint. It does not forbid complexity outright. Instead, it makes complexity costly.
6.1.7.3. L2 Regularization (Ridge): Shrinking Weights#
Ridge regression introduces an L2 penalty, which is the sum of squared coefficients.
[ \text{Loss}{\text{Ridge}} = \text{MSE} + \alpha \sum{i=1}^{n} w_i^2 ]
Because the penalty squares the coefficients, larger values are penalized more heavily. The effect is smooth shrinkage: coefficients move toward zero but rarely become exactly zero.
Intuition#
Large coefficients become expensive.
All coefficients shrink proportionally.
Variance decreases.
Overfitting is reduced.
The next example demonstrates how increasing α affects both coefficients and error.
from sklearn.linear_model import Ridge
# Generate synthetic data
np.random.seed(42)
n_samples, n_features = 50, 20
X = np.random.randn(n_samples, n_features)
true_coef = np.zeros(n_features)
true_coef[:3] = [5, -3, 2]
y = X @ true_coef + np.random.randn(n_samples) * 2
# Split data
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=42)
alphas = [0, 0.1, 1, 10, 100]
models = {}
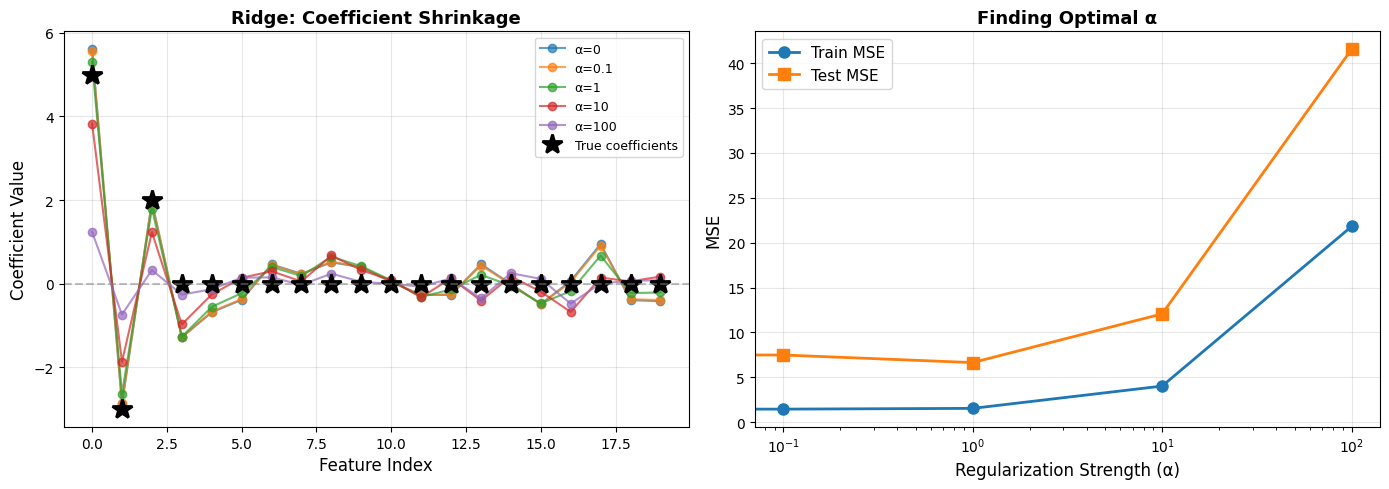
From the experiment:
Key Observations:
α = 0 produces large coefficients and overfitting.
Small α introduces mild shrinkage.
Moderate α achieves the best test performance.
Large α forces all coefficients near zero, causing underfitting.
Optimal α for this data: ~1
Ridge regression is particularly useful when features are correlated. Instead of eliminating variables, it distributes weight across them more conservatively.
Tip
Use Ridge (L2) when:
You have many correlated features
You want to keep all features but reduce their impact
You have more features than samples
Features are on different scales (combine with StandardScaler!)
6.1.7.4. L1 Regularization (Lasso): Sparse Solutions#
Lasso regression replaces the squared penalty with an absolute value penalty:
[ \text{Loss}{\text{Lasso}} = \text{MSE} + \alpha \sum{i=1}^{n} |w_i| ]
This small mathematical change has a dramatic effect. The L1 penalty encourages exact zeros in the coefficient vector.
Intuition#
Any nonzero coefficient incurs a cost.
Many coefficients collapse to exactly zero.
The model performs automatic feature selection.
In contrast to Ridge, which shrinks everything smoothly, Lasso simplifies the model structurally.
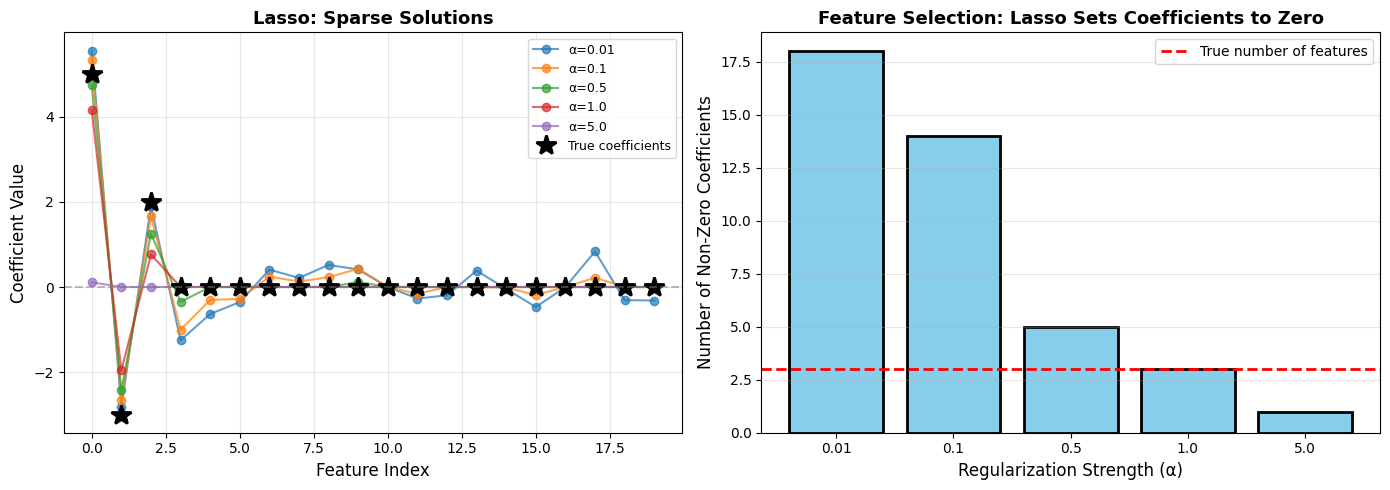
Lasso Feature Selection:
α=0.01: Selected features: [0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 11, 12, 13, 14, 15, 17, 18, 19], Number: 18/20
α=0.1: Selected features: [0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 11, 14, 15, 17], Number: 14/20
α=0.5: Selected features: [0, 1, 2, 3, 9], Number: 5/20 ✓ Correctly identified first 3 features!
α=1.0: Selected features: [0, 1, 2], Number: 3/20
α=5.0: Selected features: [0], Number: 1/20
Lasso is especially attractive when interpretability matters. A sparse model is easier to understand and communicate.
6.1.7.5. Ridge vs Lasso#
When comparing Ridge and Lasso directly, their philosophical difference becomes clear.
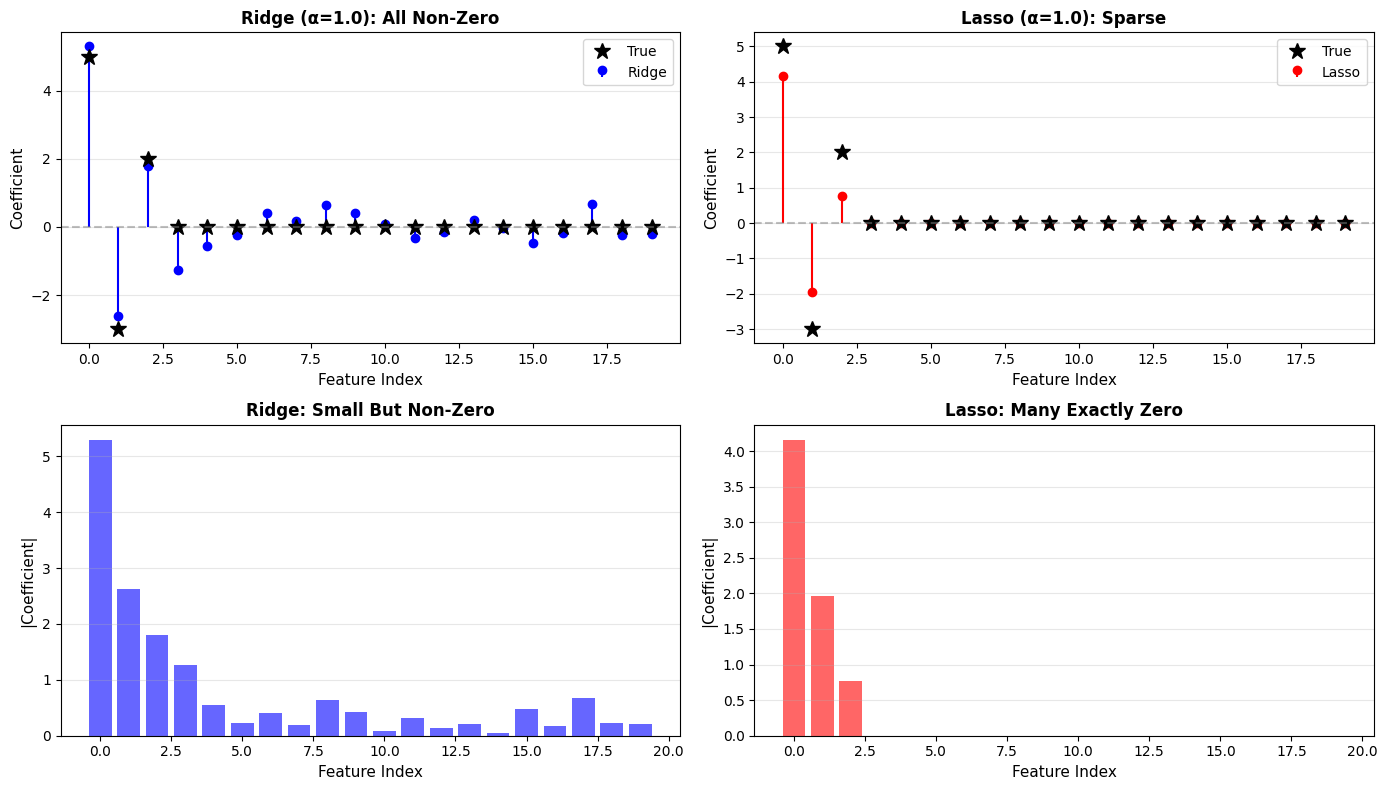
Ridge vs Lasso:
Ridge (L2):
Non-zero coefficients: 20/20
Max coefficient: 5.296
Test MSE: 6.634
Lasso (L1):
Non-zero coefficients: 3/20
Max coefficient: 4.161
Test MSE: 11.113
Aspect |
Ridge (L2) |
Lasso (L1) |
|---|---|---|
Penalty |
Sum of squared coefficients |
Sum of absolute coefficients |
Effect on coefficients |
Shrinks all toward zero |
Sets many to exactly zero |
Feature selection |
No |
Yes (automatic) |
When to use |
Many correlated features |
Suspect many features irrelevant |
Interpretability |
All features contribute |
Sparse, easy to interpret |
Computational |
Has closed-form solution |
Requires iterative optimization |
Ridge preserves all features with reduced magnitude. Lasso removes many features entirely.
The choice depends on whether you value stability across correlated features or sparsity and interpretability.
6.1.7.6. Elastic Net: Best of Both Worlds#
Elastic Net combines both penalties:
[ \text{Loss}_{\text{ElasticNet}} = \text{MSE} + \alpha \rho \sum |w_i| + \alpha (1-\rho) \sum w_i^2 ]
Here:
α controls overall regularization strength.
ρ controls the balance between L1 and L2.
When ρ = 0, the model behaves like Ridge. When ρ = 1, it behaves like Lasso.
Elastic Net is particularly useful when features are highly correlated and sparsity is still desirable. It tends to be more stable than pure Lasso while still producing simpler models.
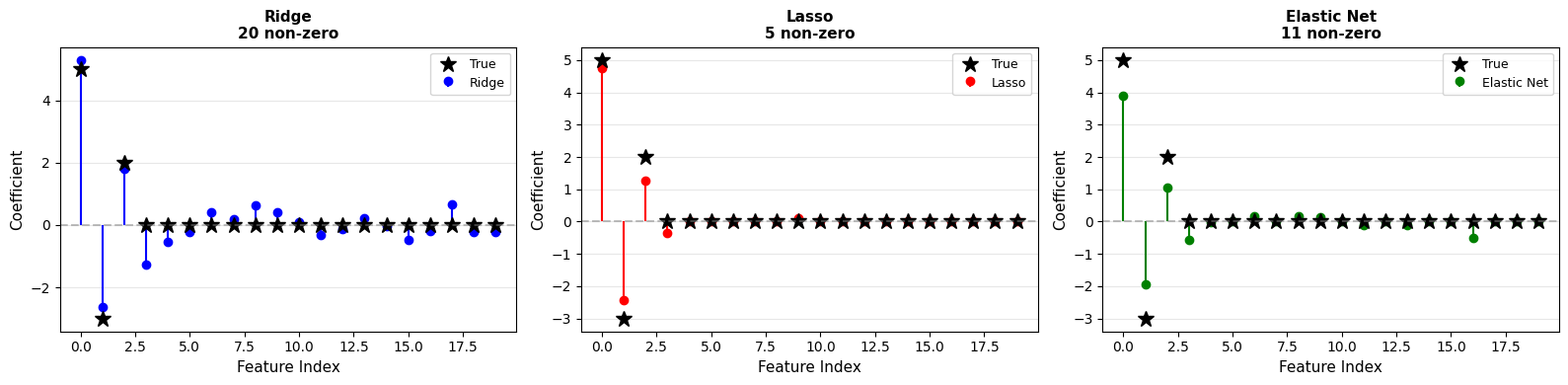
Results:
Ridge: np.int64(20) non-zero coefficients
Lasso: np.int64(5) non-zero coefficients
Elastic Net: np.int64(11) non-zero coefficients
Elastic Net balances sparsity and stability
6.1.7.7. Choosing the Regularization Strength α#
The penalty form matters, but the strength of the penalty often matters even more.
α = 0 means no regularization.
Small α introduces mild control.
Large α forces simplicity and risks underfitting.
Selecting α should not rely on guesswork. Cross-validation provides a systematic approach.
How to find the optimal α? Cross-validation!
LassoCV(alphas=array([1.00000000e-03, 1.32571137e-03, 1.75751062e-03, 2.32995181e-03,
3.08884360e-03, 4.09491506e-03, 5.42867544e-03, 7.19685673e-03,
9.54095476e-03, 1.26485522e-02, 1.67683294e-02, 2.22299648e-02,
2.94705170e-02, 3.90693994e-02, 5.17947468e-02, 6.86648845e-02,
9.10298178e-02, 1.20679264e-01, 1.59985872e-01, 2.12095089e-01,
2.81176870e-01, 3.72759372e-0...
2.68269580e+00, 3.55648031e+00, 4.71486636e+00, 6.25055193e+00,
8.28642773e+00, 1.09854114e+01, 1.45634848e+01, 1.93069773e+01,
2.55954792e+01, 3.39322177e+01, 4.49843267e+01, 5.96362332e+01,
7.90604321e+01, 1.04811313e+02, 1.38949549e+02, 1.84206997e+02,
2.44205309e+02, 3.23745754e+02, 4.29193426e+02, 5.68986603e+02,
7.54312006e+02, 1.00000000e+03]),
cv=5, max_iter=10000)In a Jupyter environment, please rerun this cell to show the HTML representation or trust the notebook. On GitHub, the HTML representation is unable to render, please try loading this page with nbviewer.org.
Parameters
| eps | 0.001 | |
| n_alphas | 'deprecated' | |
| alphas | array([1.0000...00000000e+03]) | |
| fit_intercept | True | |
| precompute | 'auto' | |
| max_iter | 10000 | |
| tol | 0.0001 | |
| copy_X | True | |
| cv | 5 | |
| verbose | False | |
| n_jobs | None | |
| positive | False | |
| random_state | None | |
| selection | 'cyclic' |
Finding Optimal Regularization Strength
Ridge best α: 2.6827
Lasso best α: 0.2812
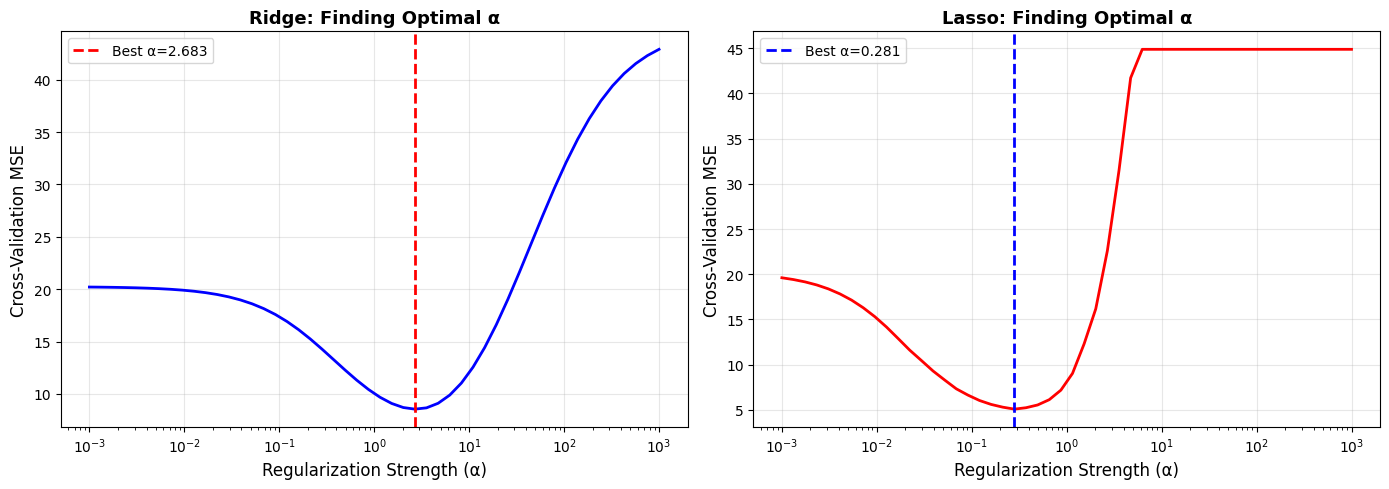
Cross-validation curves make the tradeoff visible: too little regularization increases variance, too much increases bias. The minimum point balances both.
Optimal models found via cross-validation:
Ridge test MSE: 6.989
Lasso test MSE: 5.981
Tip
Best Practice for Choosing α:
Start with a wide logarithmic range.
Use cross-validation tools such as RidgeCV or LassoCV.
For Elastic Net, tune both α and l1_ratio.
Visualize the validation curve.
Evaluate the final choice on a held-out test set.
6.1.7.8. Beyond Linear Models#
Regularization is not limited to linear regression. The principle of controlling complexity appears across machine learning.
Dropout#
In neural networks, dropout randomly disables neurons during training. This prevents units from becoming overly dependent on one another and improves generalization.
Dropout: Randomly “drop” (set to zero) neurons during training
Effect: Prevents co-adaptation of neurons, reduces overfitting
## Conceptual example (PyTorch style)
import torch.nn as nn
model = nn.Sequential(
nn.Linear(100, 50),
nn.ReLU(),
nn.Dropout(p=0.5), ## Drop 50% of neurons
nn.Linear(50, 10)
)
Early Stopping#
Another form of regularization does not modify the objective function at all. Instead, it limits training duration.
As training proceeds, training error often decreases steadily. Validation error, however, eventually begins to increase. Stopping at the minimum validation error prevents the model from overfitting.
Early Stopping: Stop training when validation error stops improving
Effect: Prevents model from overfitting by not training too long
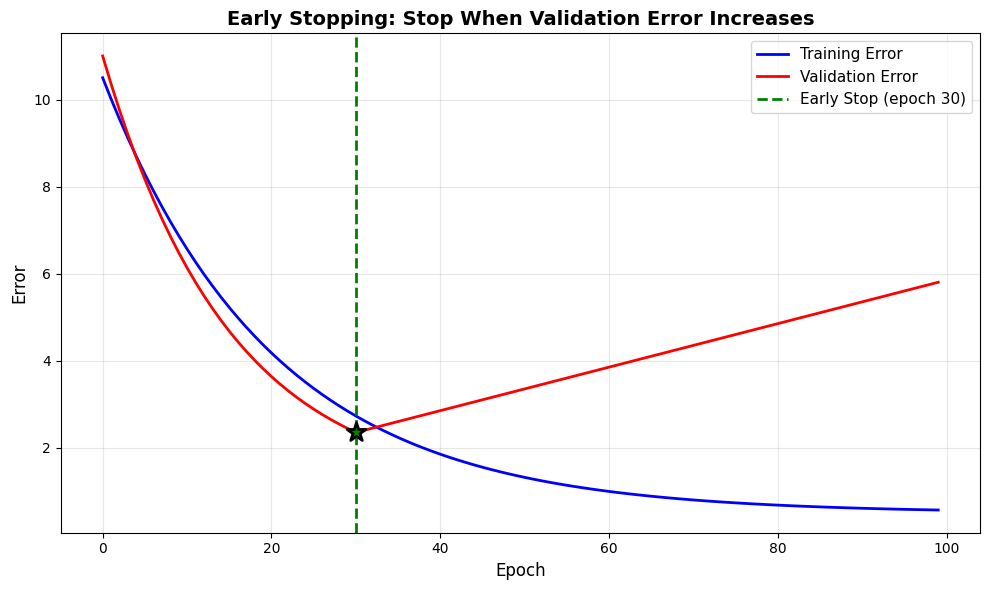
Early Stopping:
Best validation error at epoch: np.int64(30)
If we continued to epoch 100:
Training error would be: 0.57 (keeps improving)
Validation error would be: 5.80 (gets worse!)
Early stopping prevents overfitting without modifying the model
Data Augmentation#
Data augmentation increases the effective size of the dataset by generating new examples from existing ones. By exposing the model to more variation, it reduces overfitting without changing the model structure.
Examples include:
Image transformations such as rotation and flipping
Text transformations such as synonym replacement
Time series perturbations such as noise injection
Effect: Increases effective dataset size, reduces overfitting
Examples:
Images: Rotate, flip, crop, color jitter
Text: Synonym replacement, back-translation
Time series: Time warping, noise injection
6.1.7.9. Practical Guidelines#
Note
When to Use Which Regularization:
Use Ridge (L2) when:
All features may be relevant
Features are correlated
Smooth shrinkage is desired
Use Lasso (L1) when:
Many features are likely irrelevant
Interpretability is important
Automatic feature selection is needed
Use Elastic Net when:
You want both stability and sparsity
Correlated feature groups exist
Features outnumber samples
Use Dropout when:
Training deep neural networks
Use Early Stopping when:
Training iterative models
Validation error begins to increase
Computational resources are limited

