6.3.1.4. Cluster Evaluation: Measuring Quality#
You have learned how to perform clustering using K-Means Clustering, Hierarchical Clustering, and DBSCAN: Density-Based Clustering methods. The natural next question is: how do we know whether the clusters we obtained are actually good?
Unlike supervised learning, clustering does not usually come with ground truth labels. There is no direct accuracy measure. Instead, we must evaluate clusters indirectly by measuring their internal structure.
A good clustering should satisfy two key properties:
Cohesion: Points within the same cluster should be similar to one another.
Separation: Points in different clusters should be clearly distinct.
Cluster evaluation metrics are designed to quantify these ideas.
The Challenge: No Ground Truth#
In supervised learning, we compare predictions against known labels. In clustering, we typically have no such labels. This makes evaluation inherently more subtle.
There are two broad categories of evaluation metrics:
1. Internal Metrics#
These rely only on the feature data (X). They assess how well the clustering structure fits the data itself.
Silhouette Score
Davies-Bouldin Index
Calinski-Harabasz Index
2. External Metrics#
These require true labels. They are used when ground truth is available for validation purposes.
Adjusted Rand Index (ARI)
Normalized Mutual Information (NMI)
In most real applications, internal metrics are the primary tools, since labels are rarely available.
Silhouette Score: The Most Widely Used Metric#
The Silhouette Score evaluates how well each point fits within its assigned cluster compared to other clusters.
For each point (i):
a(i): Average distance to other points in the same cluster
b(i): Average distance to points in the nearest neighboring cluster
s(i) = \(\frac{b(i) - a(i)}{\max(a(i), b(i))}\)
Interpretation of (s(i)):
+1: The point is well matched to its own cluster and far from others
0: The point lies near a cluster boundary
-1: The point may be assigned to the wrong cluster
The overall Silhouette Score is simply the average of (s(i)) across all data points.
The following experiment compares silhouette scores for different values of (k).
# Try different k values
k_values = [2, 3, 4, 5]
fig, axes = plt.subplots(2, 2, figsize=(14, 12))
axes = axes.ravel()
for idx, k in enumerate(k_values):
kmeans = KMeans(n_clusters=k, init='k-means++', n_init=10, random_state=42)
labels = kmeans.fit_predict(X_sil)
# Compute silhouette
sil_score = silhouette_score(X_sil, labels)
sil_samples = silhouette_samples(X_sil, labels)
# Plot clusters
colors = ['red', 'blue', 'green', 'orange', 'purple']
for cluster_id in range(k):
cluster_mask = labels == cluster_id
axes[idx].scatter(X_sil[cluster_mask, 0], X_sil[cluster_mask, 1],
s=60, c=colors[cluster_id], alpha=0.6,
edgecolors='black', linewidths=0.5,
label=f'C{cluster_id} (sil={sil_samples[cluster_mask].mean():.2f})')
axes[idx].scatter(kmeans.cluster_centers_[:, 0],
kmeans.cluster_centers_[:, 1],
s=300, c='yellow', marker='*',
edgecolors='black', linewidths=2)
axes[idx].set_xlabel('Feature 1', fontsize=11)
axes[idx].set_ylabel('Feature 2', fontsize=11)
axes[idx].set_title(f'k={k}\nSilhouette Score: {sil_score:.3f}',
fontsize=12, fontweight='bold')
axes[idx].legend(loc='best', fontsize=8)
axes[idx].grid(True, alpha=0.3)
plt.tight_layout()
plt.show()
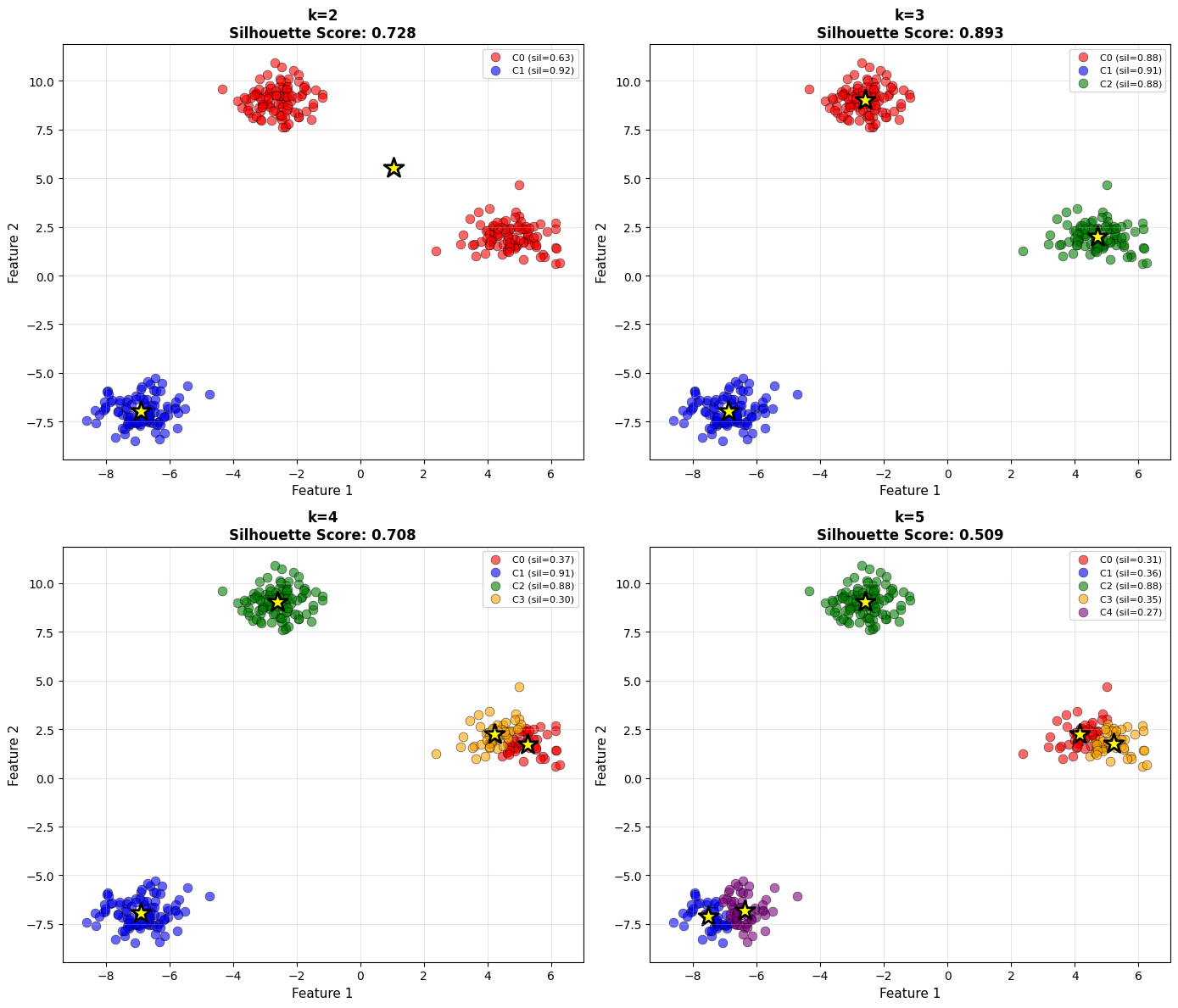
As you examine the plots, notice how both cluster structure and silhouette score change with (k). The optimal number of clusters typically corresponds to a high average silhouette value and visually well-separated groups.
Davies-Bouldin Index: Measuring Cluster Similarity#
The Davies-Bouldin Index quantifies how similar each cluster is to its most similar neighboring cluster.
It compares within-cluster scatter to between-cluster separation.
It penalizes clusters that overlap or are poorly separated.
Lower values indicate better clustering.
A value close to 0 represents strong separation.
Unlike the silhouette score, which is bounded between -1 and 1, the Davies-Bouldin Index has no fixed upper bound. Interpretation relies on relative comparison across models.
# Compare different k values
k_range = range(2, 8)
db_scores = []
sil_scores = []
for k in k_range:
kmeans = KMeans(n_clusters=k, init='k-means++', n_init=10, random_state=42)
labels = kmeans.fit_predict(X_sil)
db_score = davies_bouldin_score(X_sil, labels)
sil_score = silhouette_score(X_sil, labels)
db_scores.append(db_score)
sil_scores.append(sil_score)
# Plot
fig, axes = plt.subplots(1, 2, figsize=(14, 5))
# Davies-Bouldin (lower better)
axes[0].plot(k_range, db_scores, 'o-', linewidth=2, markersize=8, color='red')
axes[0].axvline(x=3, color='green', linestyle='--', linewidth=2,
label='True k=3')
axes[0].set_xlabel('Number of Clusters (k)', fontsize=12)
axes[0].set_ylabel('Davies-Bouldin Index', fontsize=12)
axes[0].set_title('Davies-Bouldin Index\n(Lower is Better)',
fontsize=13, fontweight='bold')
axes[0].legend()
axes[0].grid(True, alpha=0.3)
# Silhouette (higher better)
axes[1].plot(k_range, sil_scores, 'o-', linewidth=2, markersize=8, color='blue')
axes[1].axvline(x=3, color='green', linestyle='--', linewidth=2,
label='True k=3')
axes[1].set_xlabel('Number of Clusters (k)', fontsize=12)
axes[1].set_ylabel('Silhouette Score', fontsize=12)
axes[1].set_title('Silhouette Score\n(Higher is Better)',
fontsize=13, fontweight='bold')
axes[1].legend()
axes[1].grid(True, alpha=0.3)
plt.tight_layout()
plt.show()
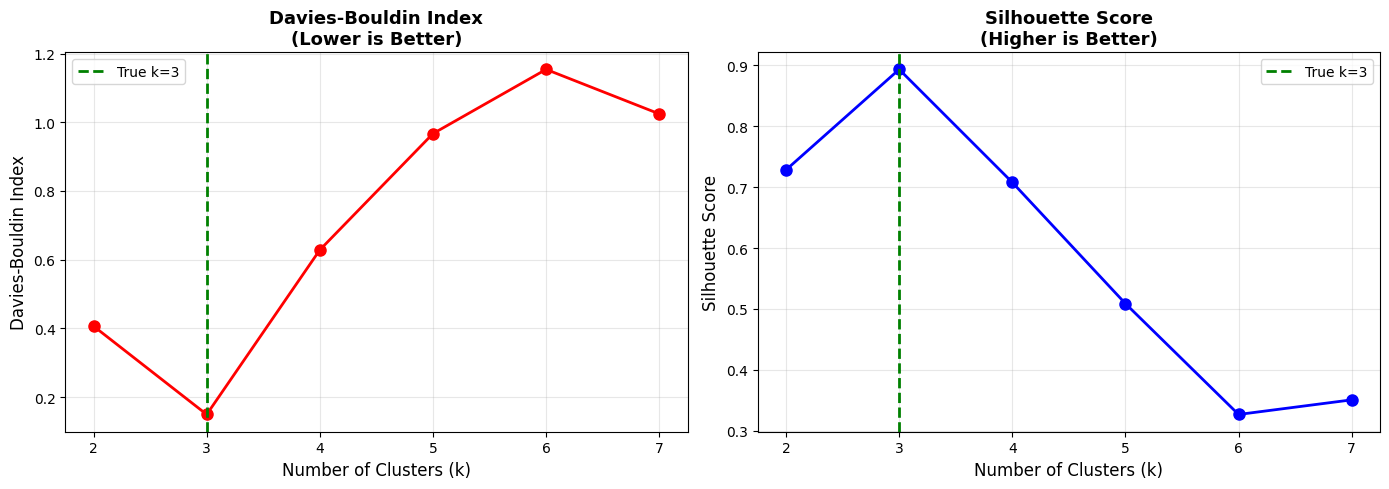
When comparing different values of (k), look for the configuration that minimizes the Davies-Bouldin score while maintaining interpretability.
Calinski-Harabasz Index: The Variance Ratio#
The Calinski-Harabasz Index, also known as the Variance Ratio Criterion, evaluates the ratio of:
Between-cluster dispersion
Within-cluster dispersion
Intuitively, good clustering should maximize separation between clusters while minimizing variation inside clusters.
Higher values indicate better-defined clusters.
# Compute for different k
ch_scores = []
for k in k_range:
kmeans = KMeans(n_clusters=k, init='k-means++', n_init=10, random_state=42)
labels = kmeans.fit_predict(X_sil)
ch_score = calinski_harabasz_score(X_sil, labels)
ch_scores.append(ch_score)
# Plot all three metrics together
fig, axes = plt.subplots(1, 3, figsize=(16, 5))
# Silhouette
axes[0].plot(k_range, sil_scores, 'o-', linewidth=2, markersize=8, color='blue')
axes[0].axvline(x=3, color='red', linestyle='--', linewidth=2, label='Best k')
axes[0].set_xlabel('k', fontsize=12)
axes[0].set_ylabel('Silhouette Score', fontsize=12)
axes[0].set_title('Silhouette\n(Higher Better)', fontsize=12, fontweight='bold')
axes[0].legend()
axes[0].grid(True, alpha=0.3)
# Davies-Bouldin
axes[1].plot(k_range, db_scores, 'o-', linewidth=2, markersize=8, color='red')
axes[1].axvline(x=3, color='green', linestyle='--', linewidth=2, label='Best k')
axes[1].set_xlabel('k', fontsize=12)
axes[1].set_ylabel('Davies-Bouldin Index', fontsize=12)
axes[1].set_title('Davies-Bouldin\n(Lower Better)', fontsize=12, fontweight='bold')
axes[1].legend()
axes[1].grid(True, alpha=0.3)
# Calinski-Harabasz
axes[2].plot(k_range, ch_scores, 'o-', linewidth=2, markersize=8, color='green')
axes[2].axvline(x=3, color='red', linestyle='--', linewidth=2, label='Best k')
axes[2].set_xlabel('k', fontsize=12)
axes[2].set_ylabel('Calinski-Harabasz Score', fontsize=12)
axes[2].set_title('Calinski-Harabasz\n(Higher Better)', fontsize=12, fontweight='bold')
axes[2].legend()
axes[2].grid(True, alpha=0.3)
plt.tight_layout()
plt.show()
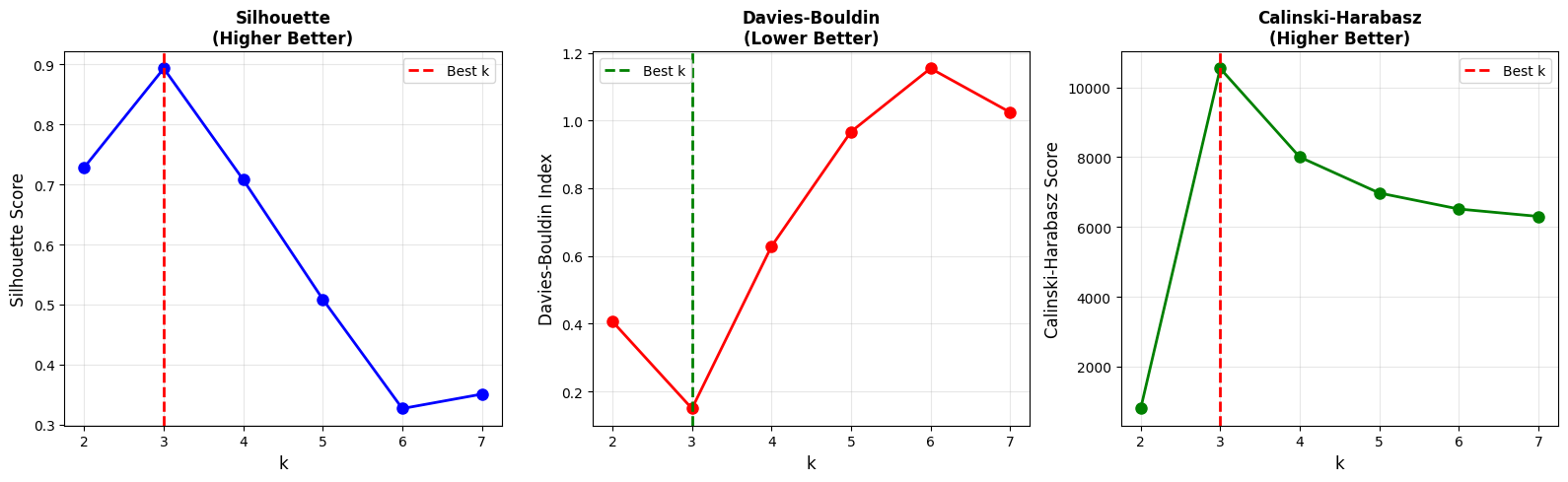
When multiple metrics agree on the same value of (k), confidence in that clustering choice increases.
Comparing Algorithms#
Evaluation metrics are also useful for comparing different clustering algorithms on the same dataset.
Here we compare:
K-Means
Hierarchical Clustering
DBSCAN
All models are applied to standardized data to ensure fair comparison. Distance-based metrics are highly sensitive to feature scale, so standardization is essential.
# Scale data
scaler = StandardScaler()
X_sil_scaled = scaler.fit_transform(X_sil)
# Try different algorithms
algorithms = {
'K-Means': KMeans(n_clusters=3, init='k-means++', n_init=10, random_state=42),
'Hierarchical': AgglomerativeClustering(n_clusters=3, linkage='ward'),
'DBSCAN': DBSCAN(eps=0.5, min_samples=5)
}
results = []
fig, axes = plt.subplots(1, 3, figsize=(16, 5))
for idx, (name, algo) in enumerate(algorithms.items()):
labels = algo.fit_predict(X_sil_scaled)
# Skip DBSCAN if only noise
n_clusters = len(set(labels)) - (1 if -1 in labels else 0)
if n_clusters > 1:
sil = silhouette_score(X_sil_scaled, labels)
db = davies_bouldin_score(X_sil_scaled, labels)
ch = calinski_harabasz_score(X_sil_scaled, labels)
else:
sil, db, ch = np.nan, np.nan, np.nan
results.append({
'Algorithm': name,
'Clusters': n_clusters,
'Silhouette': sil,
'Davies-Bouldin': db,
'Calinski-Harabasz': ch
})
# Plot
unique_labels = set(labels)
colors = ['red', 'blue', 'green', 'orange']
for k in unique_labels:
if k == -1: # Noise
class_mask = labels == k
axes[idx].scatter(X_sil_scaled[class_mask, 0],
X_sil_scaled[class_mask, 1],
s=50, c='gray', marker='x', alpha=0.5)
else:
class_mask = labels == k
axes[idx].scatter(X_sil_scaled[class_mask, 0],
X_sil_scaled[class_mask, 1],
s=50, c=colors[k % len(colors)], alpha=0.6,
edgecolors='black', linewidths=0.5)
axes[idx].set_xlabel('Feature 1 (scaled)', fontsize=11)
axes[idx].set_ylabel('Feature 2 (scaled)', fontsize=11)
if not np.isnan(sil):
axes[idx].set_title(f'{name}\nSilhouette: {sil:.3f}',
fontsize=12, fontweight='bold')
else:
axes[idx].set_title(f'{name}\n(Evaluation failed)',
fontsize=12, fontweight='bold')
axes[idx].grid(True, alpha=0.3)
plt.tight_layout()
plt.show()
results_df = pd.DataFrame(results)
display(results_df)
| Algorithm | Clusters | Silhouette | Davies-Bouldin | Calinski-Harabasz | |
|---|---|---|---|---|---|
| 0 | K-Means | 3 | 0.893395 | 0.148312 | 9577.258358 |
| 1 | Hierarchical | 3 | 0.893395 | 0.148312 | 9577.258358 |
| 2 | DBSCAN | 3 | 0.893395 | 0.148312 | 9577.258358 |
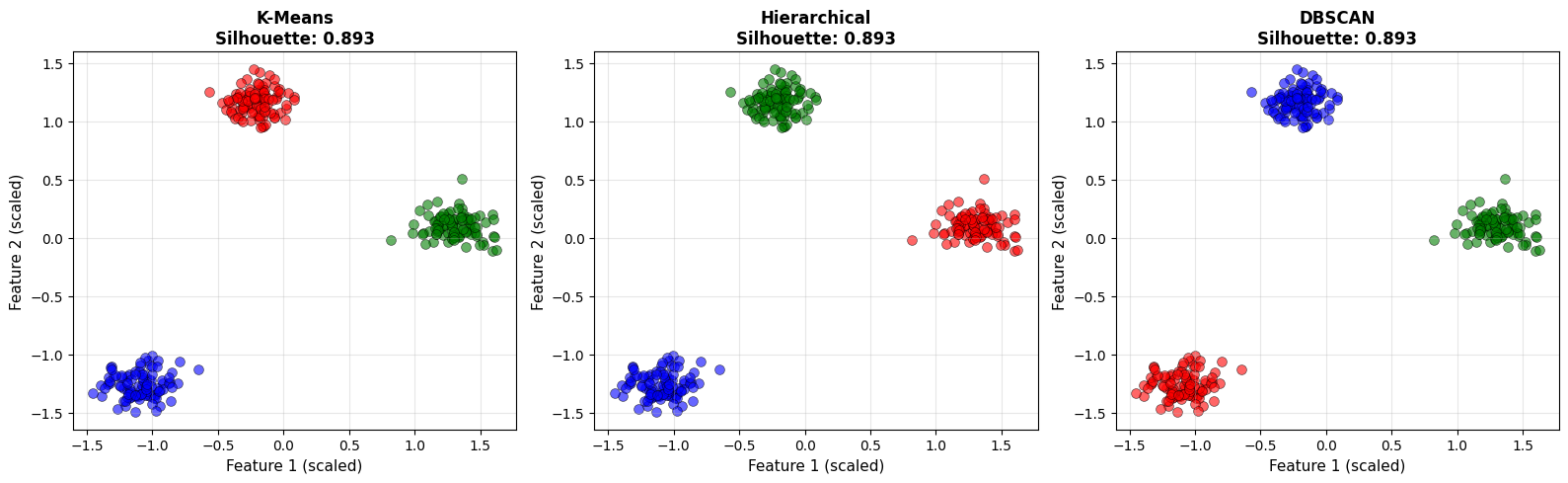
Notice that different algorithms may produce different numbers of clusters. Metrics help quantify quality, but visualization remains critical. A strong metric score does not automatically imply meaningful structure.
External Metrics: When Labels Are Available#
Occasionally, you may have ground truth labels available for validation. In such cases, external metrics provide a direct way to measure agreement between predicted clusters and true classes.
Two common external metrics:
Adjusted Rand Index (ARI)
Range: [-1, 1]
1 indicates perfect agreement
Normalized Mutual Information (NMI)
Range: [0, 1]
1 indicates perfect agreement
Unlike internal metrics, external metrics evaluate alignment with known labels rather than structural properties alone.
fig, axes = plt.subplots(1, 2, figsize=(14, 6))
for ax, labels_plot, title in zip(
axes,
[y_iris, labels_pred],
['True Species (Ground Truth)',
f'K-Means | ARI = {ari:.3f}, NMI = {nmi:.3f}']):
for cls in np.unique(labels_plot):
mask = labels_plot == cls
ax.scatter(X_iris_scaled[mask, 0], X_iris_scaled[mask, 2],
s=60, alpha=0.7, edgecolors='black', linewidths=0.5,
label=f'Class {cls}')
ax.set_xlabel('Sepal length (scaled)', fontsize=12)
ax.set_ylabel('Petal length (scaled)', fontsize=12)
ax.set_title(title, fontsize=13, fontweight='bold')
ax.legend(); ax.grid(True, alpha=0.3)
plt.tight_layout()
plt.show()
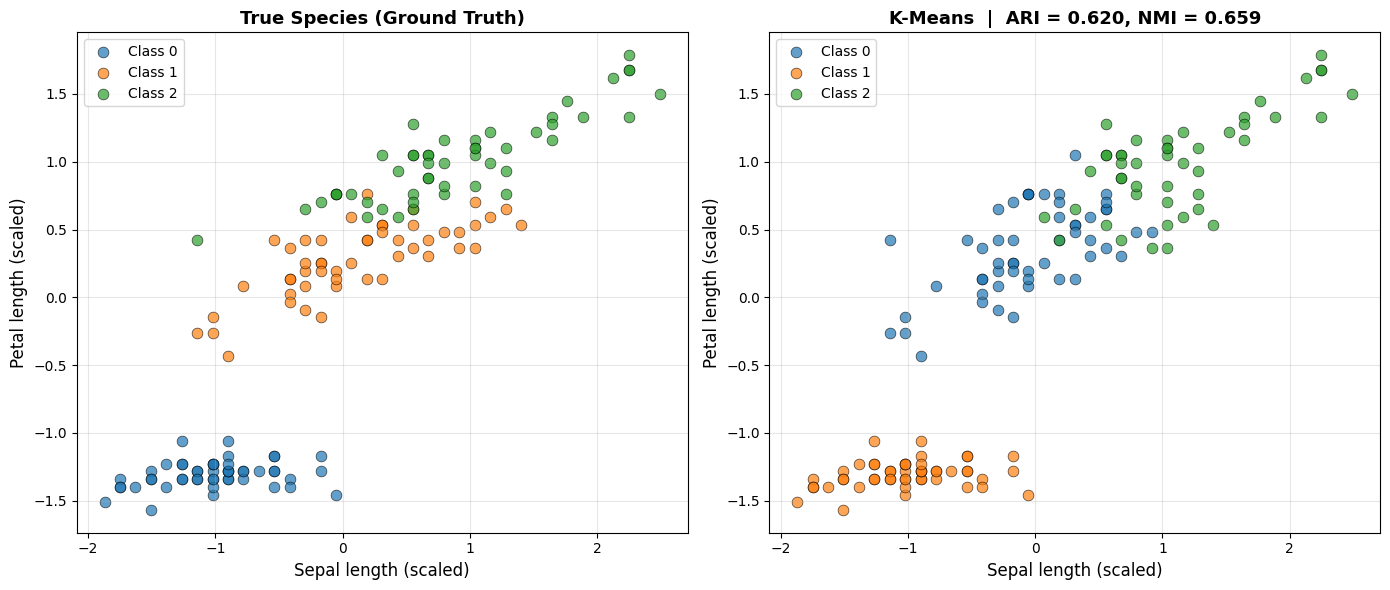
K-Means achieves an ARI of 0.62 and an NMI of 0.659, with an internal silhouette score of 0.46 and a Davies-Bouldin index of 0.834. The high ARI confirms that the discovered clusters closely align with the true species labels.
External metrics are useful for validation, but they should not be used for selecting models in purely unsupervised settings.
Practical Recommendations#
Choosing the Number of Clusters#
Use the Silhouette Score as your primary indicator. Combine it with the elbow method for K-Means when appropriate. Davies-Bouldin and Calinski-Harabasz provide complementary perspectives.
Look for agreement across multiple metrics rather than relying on a single number.
Comparing Algorithms#
Always compute metrics on the same standardized dataset.
Visualize clusters alongside metrics.
Be cautious with non-spherical or density-based structures where certain metrics may be misleading.
Metrics quantify structure, but interpretation requires context.
Silhouette Score Guidelines#
Range |
Interpretation |
|---|---|
0.70 – 1.00 |
Strong structure |
0.50 – 0.70 |
Reasonable structure |
0.25 – 0.50 |
Weak structure |
< 0.25 |
No substantial structure |
These ranges provide general intuition, not strict rules.
Common Pitfalls#
Optimizing a metric too aggressively can produce artificial solutions.
Failing to standardize features makes distance-based metrics unreliable.
High metric scores do not guarantee business relevance.
Domain knowledge often overrides small numerical differences. A cluster structure that supports meaningful decisions is more valuable than a marginal improvement in silhouette score.
Key Takeaways#
Important
Remember These Points:
Internal Metrics (No Labels)
Silhouette Score: [-1, 1], higher better
Davies-Bouldin: [0, ∞), lower better
Calinski-Harabasz: [0, ∞), higher better
External Metrics (Need Labels)
Adjusted Rand Index: [-1, 1], 1 = perfect match
Normalized Mutual Info: [0, 1], 1 = perfect match
Use for validation, not primary model selection
Silhouette Score (Most Used)
Measures cohesion and separation
Provides per-sample and overall insight
Helpful for choosing k
Practical Workflow
Compute multiple internal metrics
Look for agreement across metrics
Visualize clusters
Apply domain knowledge
Iterate
Limitations
Metrics may disagree
No single metric is perfect
Visualization remains essential
Best Practice
Use Silhouette as primary metric
Davies-Bouldin as secondary check
Always visualize results
Validate with domain experts
Cluster evaluation is not about finding a perfect score. It is about combining quantitative metrics, visual inspection, and domain understanding to arrive at meaningful structure in data.

