6.3.2.2. t-SNE#
Principal Component Analysis (PCA) finds the globally most variable directions in the data and projects everything onto them. When the interesting structure is non-linear — clusters that curve, spiral, or nest inside each other — those projections collapse the very patterns you care about.
t-SNE (t-Distributed Stochastic Neighbor Embedding) takes a fundamentally different approach: instead of preserving global variance, it preserves local neighbourhoods. Points that are close in the high-dimensional space remain close in the 2-D picture; points that are far stay far.
t-SNE produces some of the most striking data visualisations in machine learning — dense, well-separated clusters that reveal latent structure that PCA cannot see.
The Intuition#
Imagine each data point in high-D space as a person at a party. Two people are “friends” if they stand close together. t-SNE asks: can I rearrange everyone on a 2-D dance floor so that friendships are preserved? Friends should end up next to each other; strangers should remain far apart.
The key trick is that “closeness” is defined probabilistically: the probability that person \(j\) is a friend of person \(i\) falls off smoothly with distance. This soft definition is much more informative than a hard “close / not close” threshold.
The Math#
High-dimensional similarities — model closeness between \(\mathbf{x}_i\) and \(\mathbf{x}_j\) as a conditional probability using a Gaussian:
The bandwidth \(\sigma_i\) is chosen automatically so that the effective number of neighbours (the “perplexity”) matches a user-supplied target.
Symmetrise: \(p_{ij} = \tfrac{p_{j|i} + p_{i|j}}{2n}\).
Low-dimensional similarities — model closeness between embeddings \(\mathbf{y}_i\) and \(\mathbf{y}_j\) using a heavy-tailed Student-t distribution (one degree of freedom):
The heavy-tailed \(q\) distribution gives far-apart points extra room to spread out, preventing the common “crowding problem” of earlier methods.
Objective — minimise the KL divergence between the two distributions:
This is minimised with gradient descent on the positions \(\mathbf{y}\).
In scikit-learn#
from sklearn.manifold import TSNE
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
X_sc = scaler.fit_transform(X)
tsne = TSNE(n_components=2, perplexity=30, random_state=42)
X_2d = tsne.fit_transform(X_sc)
Key parameters:
Parameter |
Meaning |
|---|---|
|
Effective number of neighbours; typical range 5–50 |
|
Gradient descent steps (default 1000; increase for better convergence) |
|
Step size; |
|
Seed for reproducibility |
Warning
t-SNE is for visualisation only. The distances and orientations in the 2-D output are not meaningful in absolute terms — only the cluster structure matters. Never feed t-SNE output into a downstream ML model.
Example#
PCA vs t-SNE on the Digits Dataset#
# PCA projection
pca2 = PCA(n_components=2)
X_pca = pca2.fit_transform(X_sc)
# t-SNE projection
tsne = TSNE(n_components=2, perplexity=30, max_iter=1000, random_state=42)
X_tsne = tsne.fit_transform(X_sc)
fig, axes = plt.subplots(1, 2, figsize=(15, 6))
for ax, X_2d, method in zip(axes, [X_pca, X_tsne], ['PCA', 't-SNE']):
sc = ax.scatter(X_2d[:, 0], X_2d[:, 1], c=y,
cmap='tab10', alpha=0.7, edgecolors='k', linewidths=0.2, s=18)
ax.set_title(f'{method} Projection of Digits (64 → 2 dims)',
fontsize=13, fontweight='bold')
ax.set_xlabel('Dimension 1', fontsize=11)
ax.set_ylabel('Dimension 2', fontsize=11)
ax.grid(True, alpha=0.3)
cbar = fig.colorbar(sc, ax=axes, ticks=range(10), shrink=0.8)
cbar.set_label('Digit class', fontsize=11)
plt.suptitle('PCA vs t-SNE — Digits Dataset', fontsize=15, fontweight='bold')
plt.show()
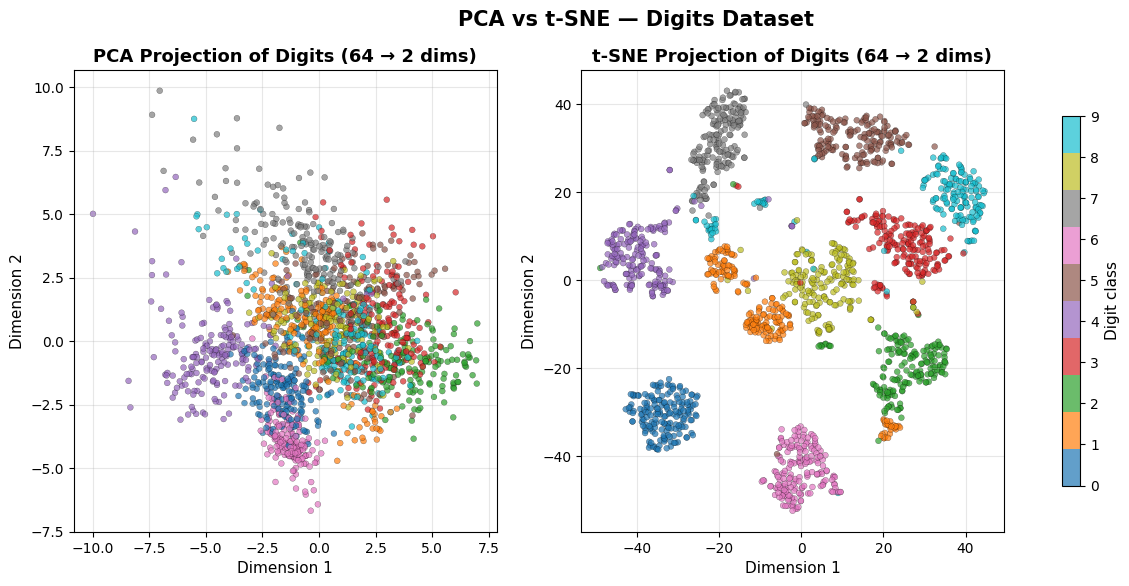
PCA shows a rough separation of digit classes but with significant overlap in the centre. t-SNE resolves all ten digit classes into cleanly separated islands, revealing a level of structure that the linear projection entirely misses.
Effect of Perplexity#
perplexities = [5, 30, 50]
fig, axes = plt.subplots(1, 3, figsize=(18, 5))
for ax, perp in zip(axes, perplexities):
emb = TSNE(n_components=2, perplexity=perp, max_iter=1000,
random_state=42).fit_transform(X_sc)
ax.scatter(emb[:, 0], emb[:, 1], c=y, cmap='tab10',
alpha=0.65, edgecolors='k', linewidths=0.15, s=16)
ax.set_title(f'Perplexity = {perp}', fontsize=12, fontweight='bold')
ax.set_xticks([]); ax.set_yticks([])
ax.grid(True, alpha=0.2)
plt.suptitle('t-SNE Perplexity Sensitivity — Digits Dataset', fontsize=14, fontweight='bold')
plt.tight_layout()
plt.show()
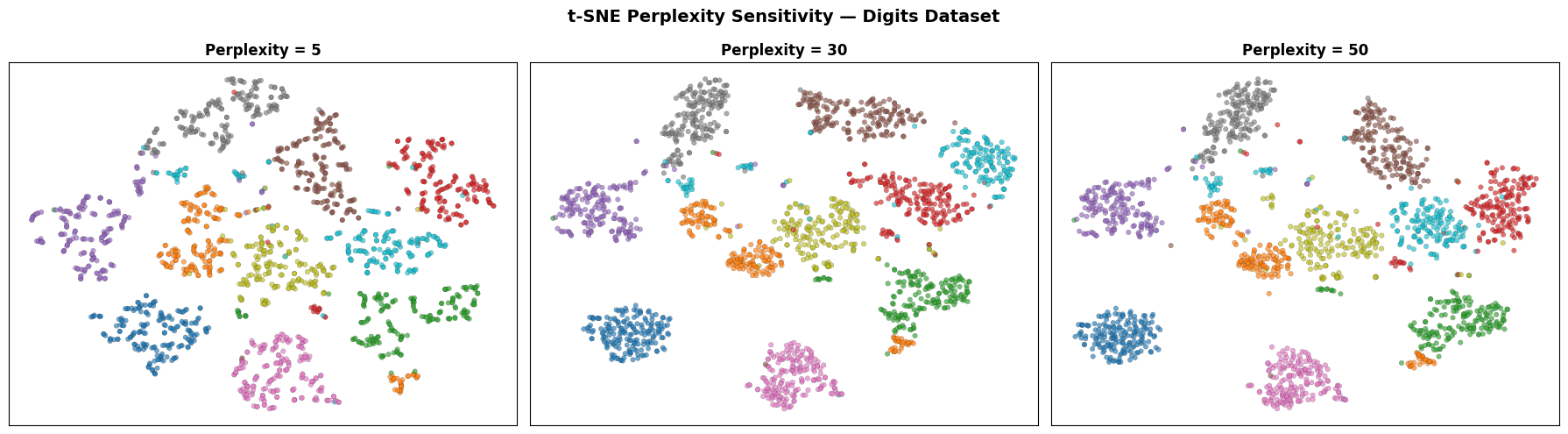
Low perplexity (5) focuses on very local structure and can fragment true clusters. High perplexity (50) brings in more global context. A value of 30 is a good starting point for most datasets.
Strengths and Weaknesses#
Strengths |
Separates clusters with remarkable clarity; handles non-linear manifolds; no assumptions about cluster shape |
Weaknesses |
Slow — \(O(n^2)\) in the basic algorithm (Barnes-Hut approximation helps); non-deterministic; distances between clusters are not interpretable; cannot be applied to new data without re-running from scratch |

